Current state and prospects of R-packages for the design of experiments
👩🏻💻 Dr. Emi Tanaka
- emi.tanaka@monash.edu
- @statsgen on Twitter
- emitanaka on GitHub
- emitanaka.org
29th June 2022
Statistical Society of Australia Canberra Branch
Take-away messages
- Slides are available at emitanaka.org/slides/ssacanberra2022/
-
Our paper is available on arXiv:
Tanaka & Amaliah (2022) Current state and prospects of R-packages for the design of experiments. arxiv.org/abs/2206.07532 -
The
edibbleR-package v0.1.0 is on CRAN with the developmental version on GitHub at github.com/emitanaka/edibble
Which vaccines are effective against COVID-19?
What diet works best for lowering insulin?
Control High-sugar
- Randomise treatments (control or high-sugar diet) to subjects
- Measure insulin level after 4 weeks
What diet works best for lowering insulin?
Control High-sugar High-fat
- In practice, experiments can be modified in consultation
- Understanding the experimental structure requires domain expertise
- This often requires communication to form shared understanding
R-packages for the
design of experiments
📜 Tanaka & Amaliah (2022) Current state and prospects of R-packages for the design of experiments. arxiv.org/abs/2206.07532
Why R-packages?
Many statistical methods are implemented in R before other languages
There are 114 R-packages in the ExperimentalDesign CRAN task view
In contrast, Python only has a handful of packages (namely
pyDOE,pyDOE2,dexpy,experimenterandGPdoemd)Characterisation of ExperimentalDesign R-packages can inform us about the practice of experimental design, with some limitations of course!
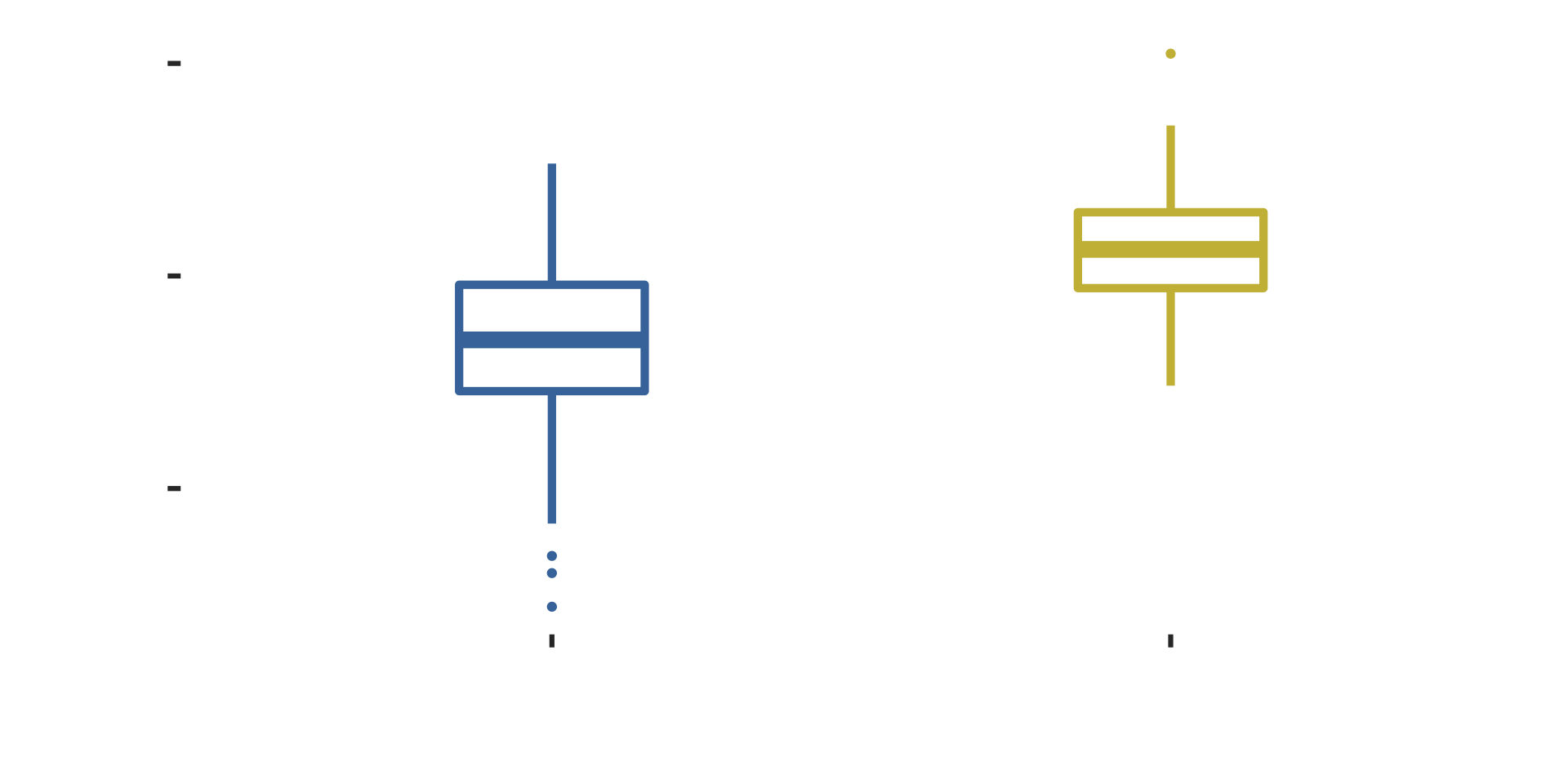
Downloads of ExperimentalDesign packages
Small number of packages dominate downloads
Lorenz curve: consider download as “wealth”
ExperimentalDesign is the least collaborative
Intra-connectivity = percentage of packages that imports, suggests or depends on other packages within the same topic
agricolae package
“Recipe”-based experimental designs
diets <- c("Control", "High-sugar", "High-fat")
# Completely Randomised Design
agricolae::design.crd(trt = diets, r = 10)
# Randomised Complete Block Design
agricolae::design.rcbd(trt = diets, r = 10)
exercise <- c("Yes", "No")
# Factorial design
agricolae::design.ab(trt = c(length(diets), length(exercise)), r = 10)
# Split-plot design
agricolae::design.split(trt1 = diets, trt2 = exercise, r = 10) AlgDesign package
“Recipe”-based optimal designs
diets <- c("Control", "High-sugar", "High-fat")
exercise <- c("Yes", "No")
data <- expand.grid(trt1 = diets, trt2 = exercise, rep = 1:10)
# Optimal design with Federov's exchange algorithm
AlgDesign::optFederov(~ trt1 + trt2 + trt1:trt2,
data = data,
nTrials = 40,
criterion = "D")
# Optimal design via Monte Carlo
AlgDesign::optMonteCarlo(~ trt1 + trt2 + trt1:trt2,
data = data,
nTrials = 40,
criterion = "D")
# Optimal design blocking
AlgDesign::optBlock(~ trt1 + trt2 + trt1:trt2,
withinData = data,
blocksizes = rep(10, 6),
criterion = "D")
# But what are the units??Good design considers units and treatments first, and then allocates treatments to units. It does not choose from a menu of named designs.
edibblestands for experimental design table (ortibble)deggustalludes to the design of experiments as ggplot objectA work-in-progress book “The Grammar of Experimental Designs” can be found at
EXPERIMENT 1

Hypothesis:
- increasing plant diversity leads to increasing soil microbial biomass and enzyme activity,
- higher temperature decreases soil microbial biomassand enzyme activity, and
- higher plant diversity buffers effects of elevated temperature on soil microbialbiomass and enzyme activity.
EXP 1 🌾 
MATERIALS AND METHODS
Experimental design
The present study took place in the BAC experiment site established at the Cedar Creek Ecosystem Science Reserve, Minnesota, USA. The site occurs on a glacial outwash plain with sandy soils. Mean temperature during the growing season (April–September) was 15.98°C in 2011 and 17.18°C in 2012. Precipitation during the growing season was 721 mm in 2011. The growing season in 2012 was considerably drier, with 545 mm rainfall.
Experimental plots (9×9 m) were planted in 1994 and 1995 with different plant communities spanning a plant diversity gradient of one, four, and 16 species, which were randomly chosen from the species listed below (Tilman et al. 2001). The grassland prairie species belonged to one of five plant functional groups: C3 grasses (Agropyron smithii Tydb., Elymus canadensis L., Koeleria cristata (Ledeb.) Schult., Poa pratensis L.), C4 grasses (Andropogon gerardii Vitman., Panicum virgatum L., Schizachyrium scoparium (Michx.) Nash, Sorghas-trum nutans (L.) Nash), legumes (Amorpha canescens Pursh., Lespedeza capitata Michx., Lupinus perennis L., Petalostemum purpureum (Vent.) Rydb., Petalostemum villosum Spreng.), nonlegume forbs (Achillea millefolium L., Asclepias tuberosa L., Liatris aspera Michx., Monarda fistulosa L., Soldidago rigida L.), and woody species (Quercus ellipsoidalis E. J. Hill, Quercus macro-carpa Michx.). The individuals of those two woody species (Quercus spp.), which were small in size and rare because of low survival, were removed from all plots in which they occurred in 2010.
In addition to the manipulation of plant diversity,the plots were divided into three subplots (2.5×3.0 m). Heat treatments were applied from March to November each year, beginning in 2009, using infrared lamps 1.8 m above ground emitting 600 W (which caused a 1.5°C increase in soil temperature for vegetation-freesoils) and 1200 W (which caused a 3°C increase; Valpine and Harte 2001, Kimball 2005, Whittingtonet al. 2013) to increase the surface soil temperature of each subplot (see Plate 1). To account for possible shading effects, metal flanges and frames were hungover control subplots. An average across all vegetated plots, temperature manipulations elevated soil temperature at 1 cm depth by 1.18°C in the low warming (+1.5°C) treatment and by 2.69°C in the high warming (+3°C) treatment, and at 10 cm depth temperature by 1.00°C in the low warming (+1.5°C) treatment and by 2.16°C in the high warming (+3°C) treatment.
Soil samples of three subplots in each of 27 experimental plots were taken; due to technical difficulties we could only analyze 66 samples out of 81 existing subplots (monoculture, 10 replicates in ambient +0°C treatment, eight replicates in +1.5°C treatment, nine replicates in +3°C treatment; four species mixture, six replicates in ambient +0°C treatment, six replicates in +1.5°C treatment, seven replicates in +3°C treatment; 16 species mixture, six replicates in ambient +0°C treatment, six replicates in +1.5°C treatment, eight replicates in +3°C treatment). The BAC plots are a representative subset of the plots in the biodiversity experiment E120 at Cedar Creek, which were assembled as random draws of a given number of species from the species pool (Zak et al. 2003). Given low heterogeneity of soil abiotic conditions at the start of the experiment, the experiment was not blocked.
EXP 1 🌾 
MATERIALS AND METHODS
Experimental design (condensed version)
- Experimental plots were planted with different plant communities spanning a plant diversity gradient of one, four, and 16 species, which were randomly chosen from the species listed (5 plant functional groups – 19 species in total)
- Plots were divided into three subplots
- Heat treatments were applied to subplots emitting 600 W which caused a 1.5°C increase in soil temperature for vegetation-free soils) and 1200 W (which caused a 3°C increase) (control with 0°C included)
- Soil samples of three subplots in each of 27 experimental plots were taken
- Given low heterogeneity of soil abiotic conditions at the start of the experiment, the experiment was not blocked.
This is in fact a split-plot design!
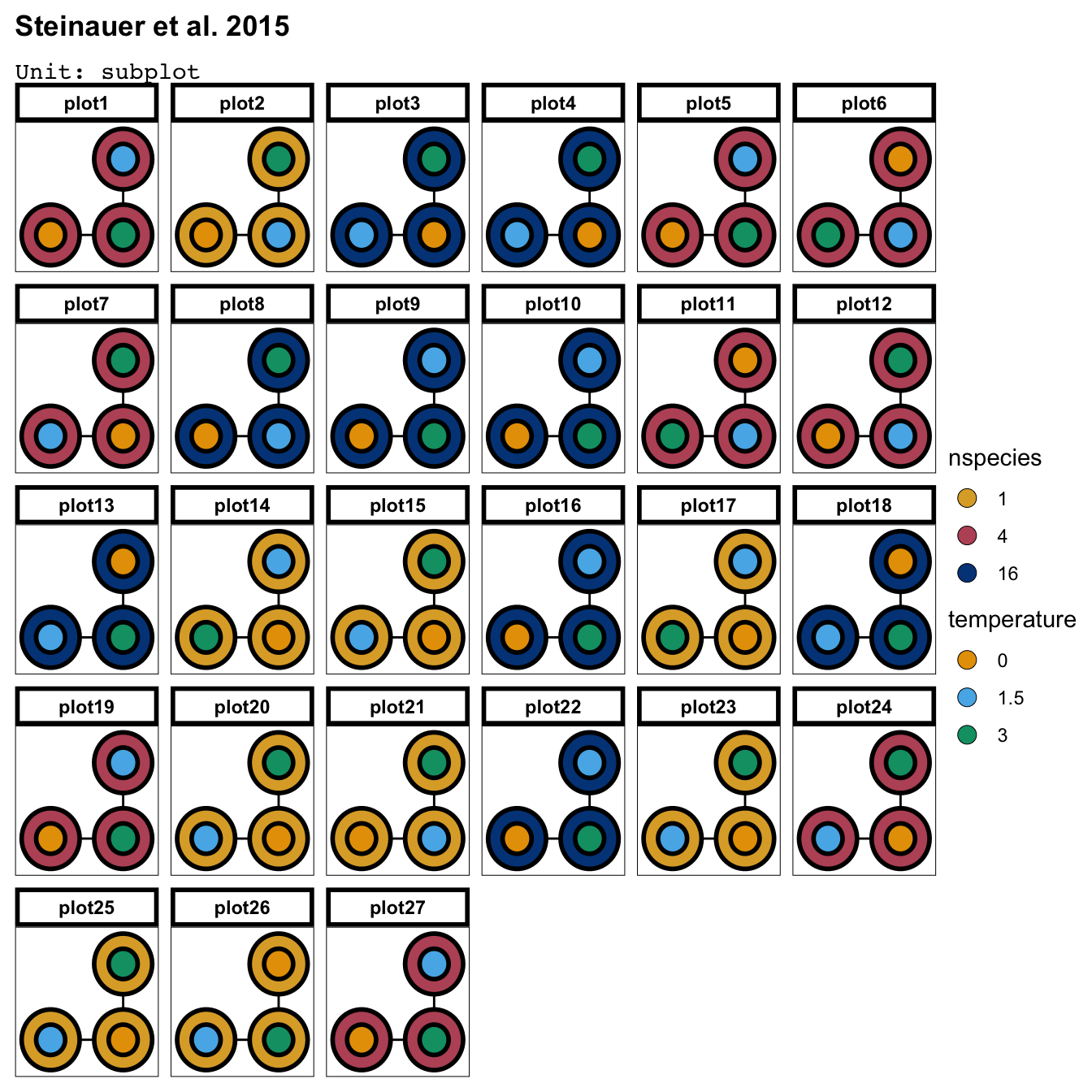
library(edibble)
des1 <- design("Steinauer et al. 2015") %>%
set_units(plot = 27,
subplot = nested_in(plot, 3)) %>%
set_trts(nspecies = c(1, 4, 16),
temperature = c(0, 1.5, 3)) %>%
allot_trts( nspecies ~ plot,
temperature ~ subplot) %>%
assign_trts("random") %>%
serve_table()
des1# Steinauer et al. 2015
# An edibble: 81 x 4
plot subplot nspecies temperature
<unit(27)> <unit(81)> <trt(3)> <trt(3)>
1 plot1 subplot1 4 0
2 plot1 subplot2 4 3
3 plot1 subplot3 4 1.5
4 plot2 subplot4 1 0
5 plot2 subplot5 1 1.5
6 plot2 subplot6 1 3
7 plot3 subplot7 16 1.5
8 plot3 subplot8 16 0
9 plot3 subplot9 16 3
10 plot4 subplot10 16 1.5
# … with 71 more rowsWhy edibble?
- Semantics of the syntactic sugar are aligned with basic terms in experimental design (i.e. code can be used for communication)
- User intention is better captured
- Experimental structure are built up step-by-step
- an incomplete structure is allowed and stored as a graph
- easy to modify or replace a single step instead of redoing whole structure
- A number of downstream benefits…
Downstream Benefit #1 Easy-and-quick visualisation
Downstream Benefit #2 Cutomise visualisation
library(deggust)
autoplot(des1, aspect_ratio = 1.5) +
# the result is a ggplot object --
# add ggplot functions on top
# as you like to customise the plot!
theme(
legend.position = "bottom",
text = element_text(size = 18),
plot.background =
element_rect(color = "black",
linetype = "dashed",
fill = "wheat"),
plot.margin = margin(20, 20, 20, 20),
plot.subtitle =
element_text(margin = margin(0, 0, 10, 0))) +
scale_fill_brewer(palette = "Reds")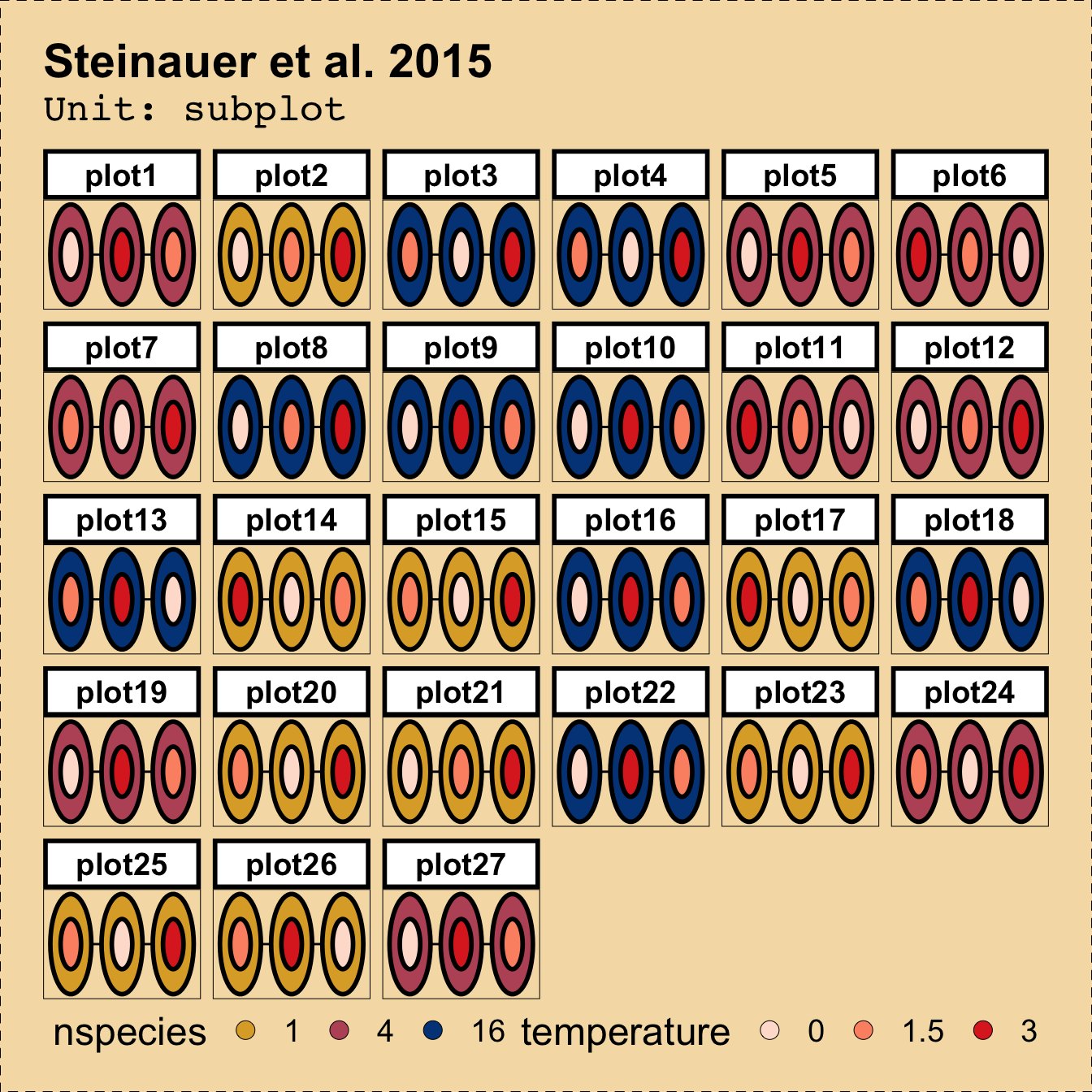
Downstream Benefit #3 Set record
- In
edibble, records are intended variables, e.g. responses, that will be measured or observed - You can set expectations of the record (plausible values) and simulate records, censoring values (default as missing) outside of expectations, or export data with data validation
library(simulate) # remotes::install_github("emitanaka/simulate")
des1 %>%
set_rcrds(microbial_biomass = subplot) %>%
expect_rcrds(microbial_biomass >= 0) %>%
simulate_rcrds(microbial_biomass = sim_normal(mean = 0.05, sd = 0.5)) # dev only# Steinauer et al. 2015
# An edibble: 81 x 5
plot subplot nspecies temperature microbial_biomass
<unit(27)> <unit(81)> <trt(3)> <trt(3)> <dbl>
1 plot1 subplot1 4 0 NA
2 plot1 subplot2 4 3 NA
3 plot1 subplot3 4 1.5 0.380
4 plot2 subplot4 1 0 NA
5 plot2 subplot5 1 1.5 0.334
6 plot2 subplot6 1 3 NA
7 plot3 subplot7 16 1.5 0.00427
8 plot3 subplot8 16 0 0.836
9 plot3 subplot9 16 3 0.0667
10 plot4 subplot10 16 1.5 0.742
# … with 71 more rowsSpecifying unbalanced designs
MATERIALS AND METHODS
Experimental design (condensed version)
- Experimental plots were planted with different plant communities spanning a plant diversity gradient of one, four, and 16 species, which were randomly chosen from the species listed (5 plant functional groups – 19 species in total)
des2 <- design("Steinauer et al. 2015 Part II") %>%
set_units(plot = 27,
plant = nested_in(plot, 1:9 ~ 1,
10:18 ~ 4,
19:27 ~ 16)) %>%
set_trts(species = 19) %>%
allot_trts(species ~ plant) %>%
assign_trts("random") %>%
serve_table()
options(deggust.nfill_max = 3)
autoplot(des2) + facet_wrap(~plot, nrow = 2)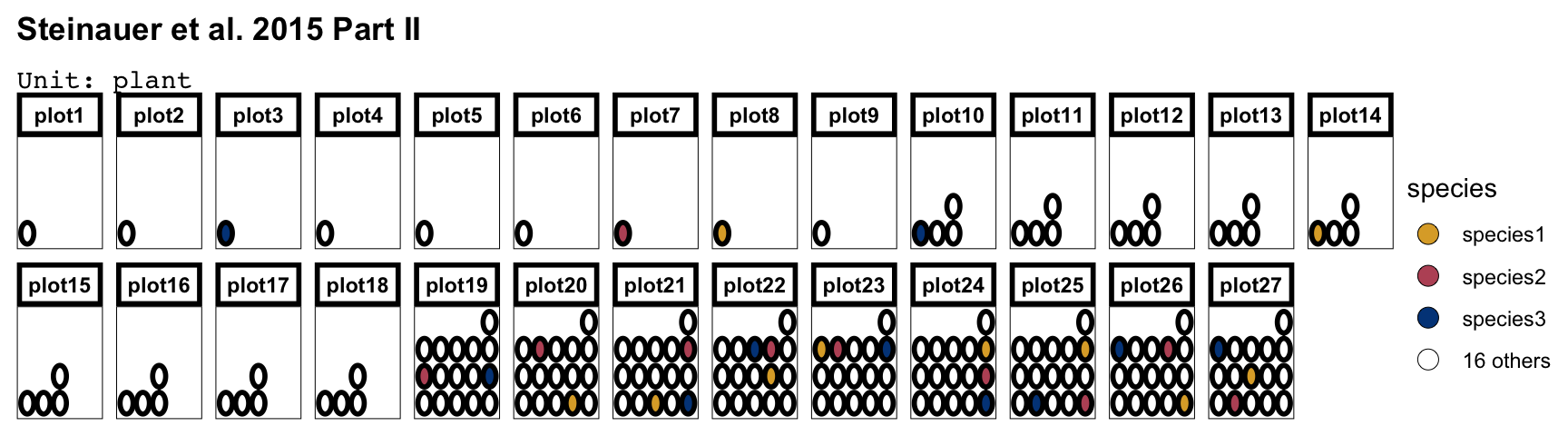
EXPERIMENT 2

Aim:
- to compare the impact of grazing, mineral fertilizer and hay cut date on the vegetation
EXP 2 🌱 
- Nine contiguous 20 x 30-m paddocks (plots), to compare the impact of grazing, mineral fertilizer and hay cut date on the vegetation in a split-split-plot design (Fig. 1).
- Three grazing treatments were randomly allocated to the three main plots in each of three blocks aligned across the 5° south facing slope of the field.
- The grazing treatments were: (i) no grazing; (ii) autumn grazing with six beef cattle and calves allowed free access to the paddocks during September and October each year; (iii) autumn grazing as in (ii) above, plus spring grazing for 1 week in early/mid May with three Swaledale ewes confined to the plots.
- Each main plot (paddock) was divided into three 10 x 20-m sub-plots and three hay cut dates (14 June, 21 July, 1 September) were randomly allocated to these.
- Each sub-plot was then divided into two 10 x 10-m sub-sub-plots and two fertilizer treatments (no fertilizer, 80 kg ha-1 nitrogen plus 40 kg ha-1 phosphorus and potassium) were randomly allocated to these.
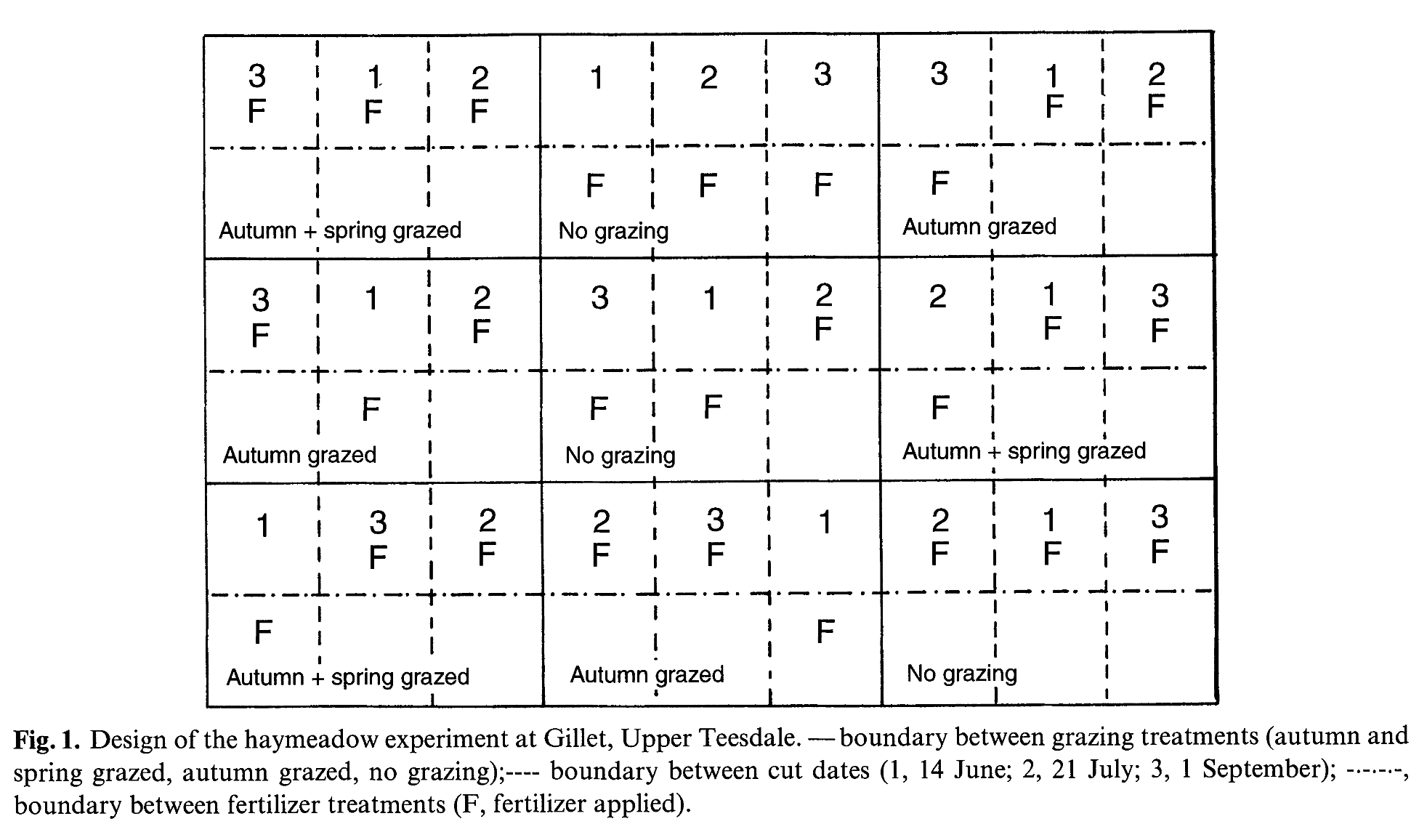
EXP 2 🌱 
- Nine contiguous 20 x 30-m paddocks (plots), to compare the impact of grazing, mineral fertilizer and hay cut date on the vegetation in a split-split-plot design (Fig. 1).
- Three grazing treatments were randomly allocated to the three main plots in each of three blocks aligned across the 5° south facing slope of the field.
- The grazing treatments were: (i) no grazing; (ii) autumn grazing with six beef cattle and calves allowed free access to the paddocks during September and October each year; (iii) autumn grazing as in (ii) above, plus spring grazing for 1 week in early/mid May with three Swaledale ewes confined to the plots.
- Each main plot (paddock) was divided into three 10 x 20-m sub-plots and three hay cut dates (14 June, 21 July, 1 September) were randomly allocated to these.
- Each sub-plot was then divided into two 10 x 10-m sub-sub-plots and two fertilizer treatments (no fertilizer, 80 kg ha-1 nitrogen plus 40 kg ha-1 phosphorus and potassium) were randomly allocated to these.
design("Smith et al. 1996") %>%
set_units(block = 3,
mainplot = nested_in(block, 3),
subplot = nested_in(mainplot, 3),
subsubplot = nested_in(subplot, 2)) %>%
set_trts(grazing = c("none", "autumn", "autumn+spring"),
cut_date = c("14 Jun", "21 Jul", "1 Sep"),
fertilizer = c("yes", "no")) %>%
# allot_table() is short hand for
# allot_trts(), assign_trts(), serve_table()
allot_table( grazing ~ mainplot,
cut_date ~ subplot,
fertilizer ~ subsubplot) # Smith et al. 1996
# An edibble: 54 x 7
block mainplot subplot subsubplot
<unit(3)> <unit(9)> <unit(27)> <unit(54)>
1 block1 mainplot1 subplot1 subsubplot1
2 block1 mainplot1 subplot1 subsubplot2
3 block1 mainplot1 subplot2 subsubplot3
4 block1 mainplot1 subplot2 subsubplot4
5 block1 mainplot1 subplot3 subsubplot5
6 block1 mainplot1 subplot3 subsubplot6
7 block1 mainplot2 subplot4 subsubplot7
8 block1 mainplot2 subplot4 subsubplot8
9 block1 mainplot2 subplot5 subsubplot9
10 block1 mainplot2 subplot5 subsubplot10
# … with 44 more rows, and 3 more variables:
# grazing <trt(3)>, cut_date <trt(3)>,
# fertilizer <trt(2)>- Getting users to clearly express, abstract and clarify the desired experimental design is a large gain across many fields
- This is only possible through our collective efforts in translating designs to a standardised, common interface/language 🤝
- Slides at emitanaka.org/slides/ssacanbera2022/
-
install.packages("edibble")
remotes::install_github("emitanaka/deggust") - Paper at arxiv.org/abs/2206.07532
- emitanaka.org/edibble-book
- emi.tanaka@monash.edu