Statistical Communication and Workflow
STAT1003 – Statistical Techniques
Dr. Emi Tanaka
Australian National University
These slides are best viewed on a modern browser like Google Chrome on a desktop or laptop. Some interactive components may require some time to fully load.
Importing and Exporting Data
Data file formats
- Data are stored as a file, which points to a block of computer memory.
- A file format signals a way to interpret the information stored in the computer memory.
- A file with the extension “csv” (comma-separated values) uses a comma as a delimiter while “tsv” uses tabs as a delimiter.
Reading and writing CSV files
File paths
- Your file has to be in the right location to be read!
- To point to the right location of the data , you may use
- a relative path (e.g.
data/data.csv) or - an absolute path (e.g.
C:\\user/myproject/data.csv)
- a relative path (e.g.
You should avoid using absolute path! Why?
- You can get and set the current path using
getwd()andsetwd(), respectively.
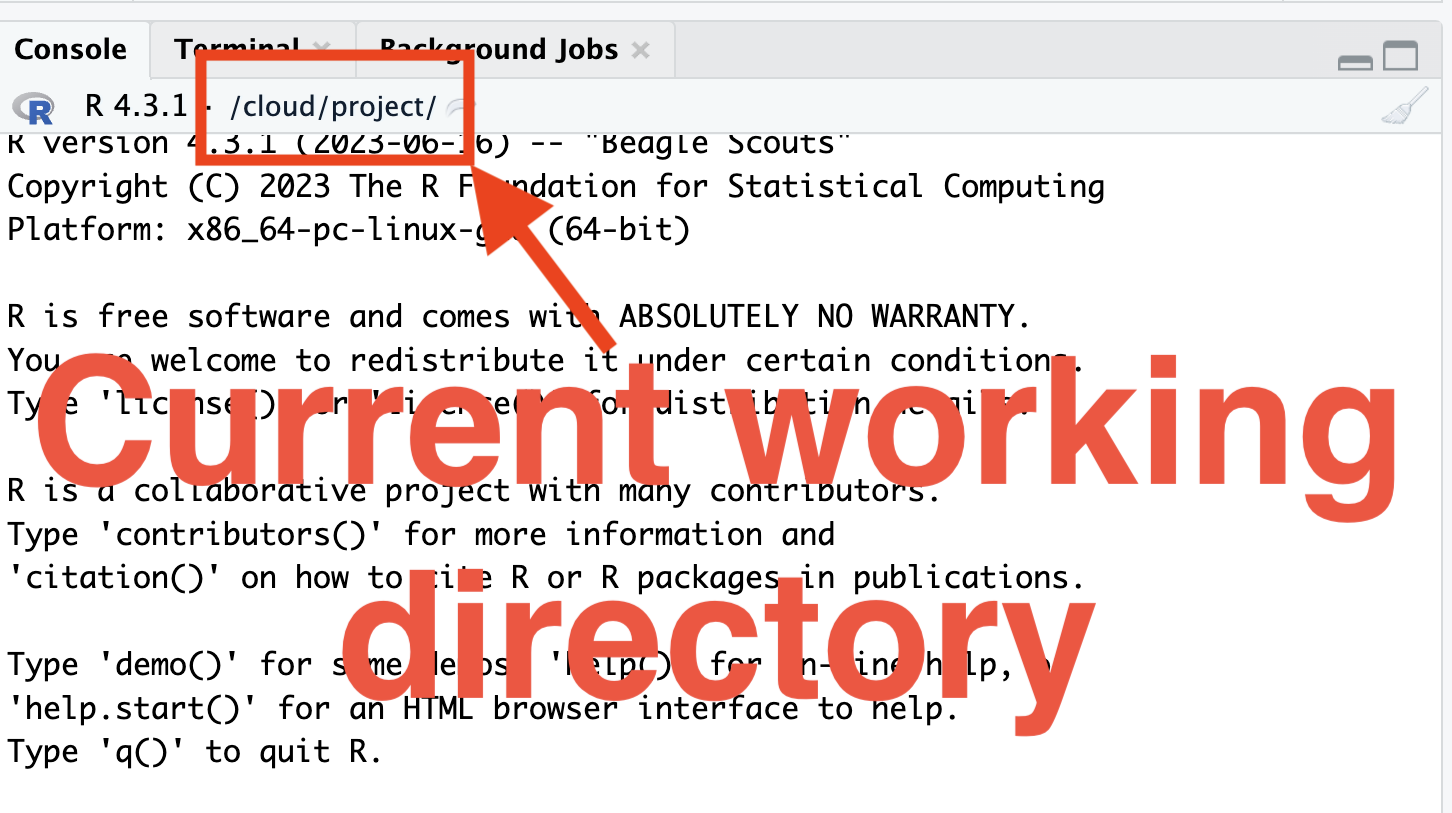
Folder structure
- Your folder structure depends on the project, but it is generally a good idea to have:
- a main project folder
- Within the main project folder, a separate folder for:
- data
- code/script/analysis
- report/paper/outputs.
- Within RStudio, you can create a project file (with an .Rproj extension).
- Double clicking on this .Rproj file launches RStudio Desktop with the current working directory set to the location of the project file.
- You can create this project file by going to RStudio > File > New Project …
Binary formats
- Data can also be stored as a binary format (e.g.
.RData,.rdaorrds). .RData,.rdaorrdssaves R objects so you don’t need the data to be in a data.frame.- However, these formats are specific to R and thus not easily portable to other software.
Reading Excel sheets
- Data can also come in a propriety format (e.g. xls and xlsx) – these require special ways to open/view/read it.
data/template_Morris.xlsx
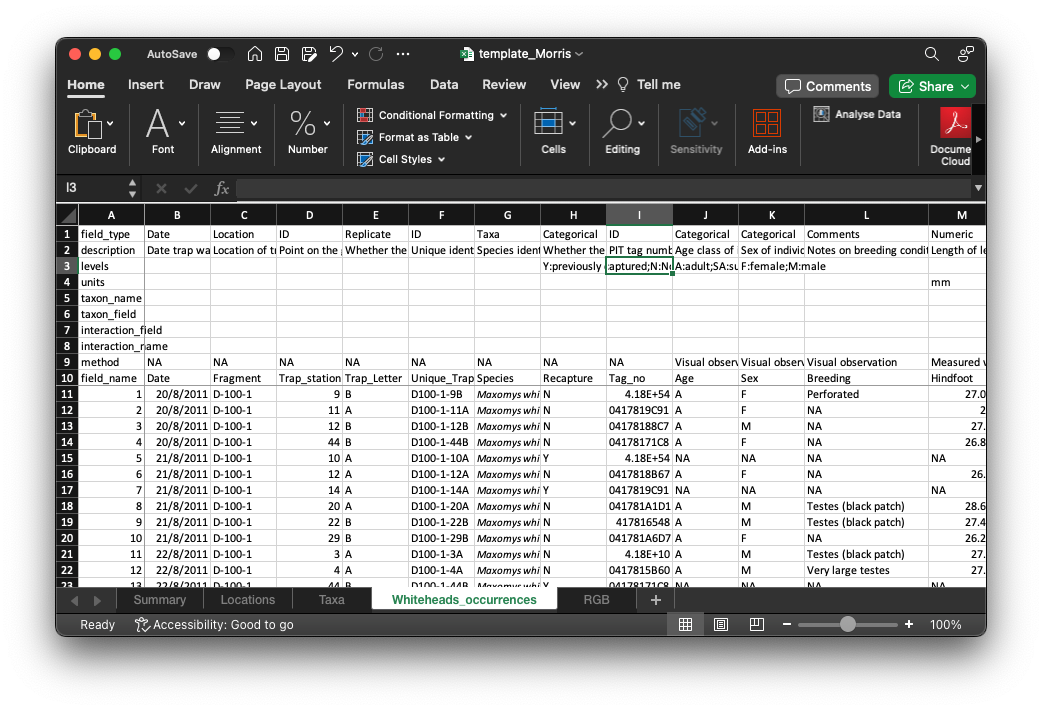
Importing through the GUI
In RStudio Desktop, you can click on the file for importing via GUI.
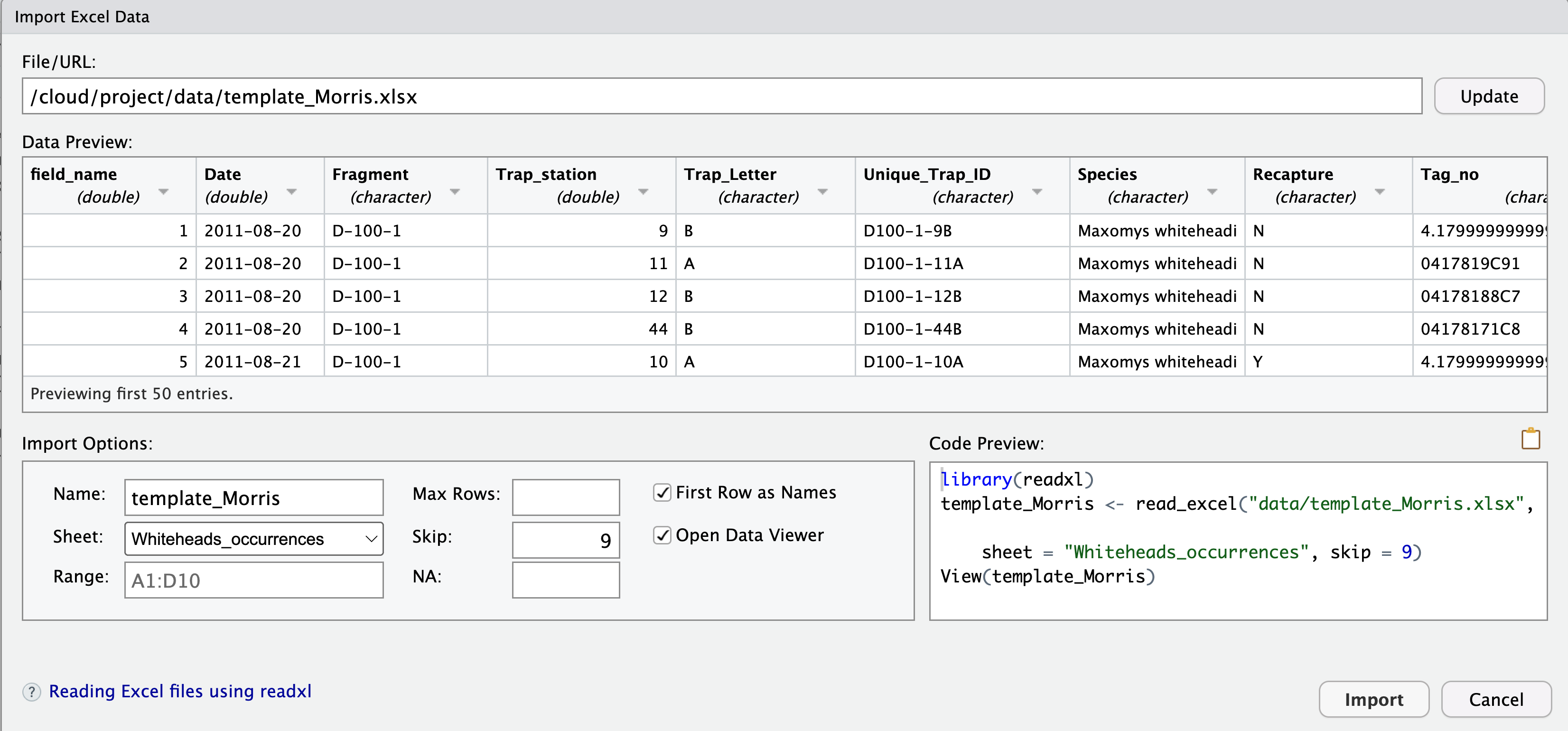
Formatting or editing data
Unless you are responsible for entering the data, you should never modify the original, stored data (note: exceptions do apply).
- For scientific integrity, any modification to the original data should be recorded in a reproducible manner (e.g. by programming in R!) so that you can trace the exact modifications.
Summary
- You can use
readr::read_csv()andreadr::write_csv()to read and write CSV files. - You can use
readxl::read_xlsx()to read Excel files. - Save a single R object using
saveRDS()(recommended) and multiple objects usingsave(). - Load R objects using
readRDS()orload(). - In RStudio Desktop, you can click on the file for importing via GUI.
- Set up R Projects and use relative path to data files.
- Don’t ever modify the raw data!
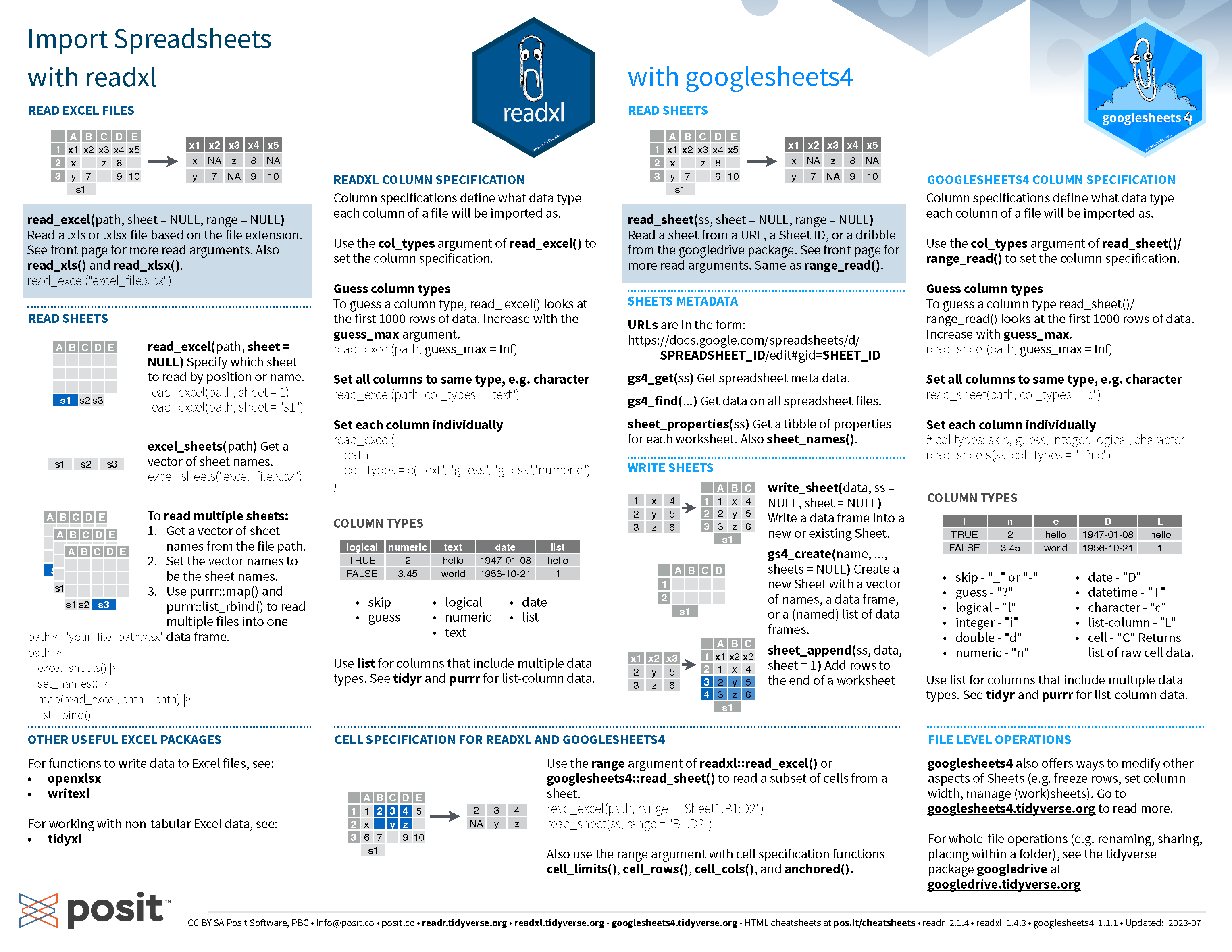
Data import cheatsheet


What is Quarto?
Quarto in a nutshell
Quarto integrates text + code in one source document with ability to render to many output formats (via Pandoc), e.g. docx, pdf or html.
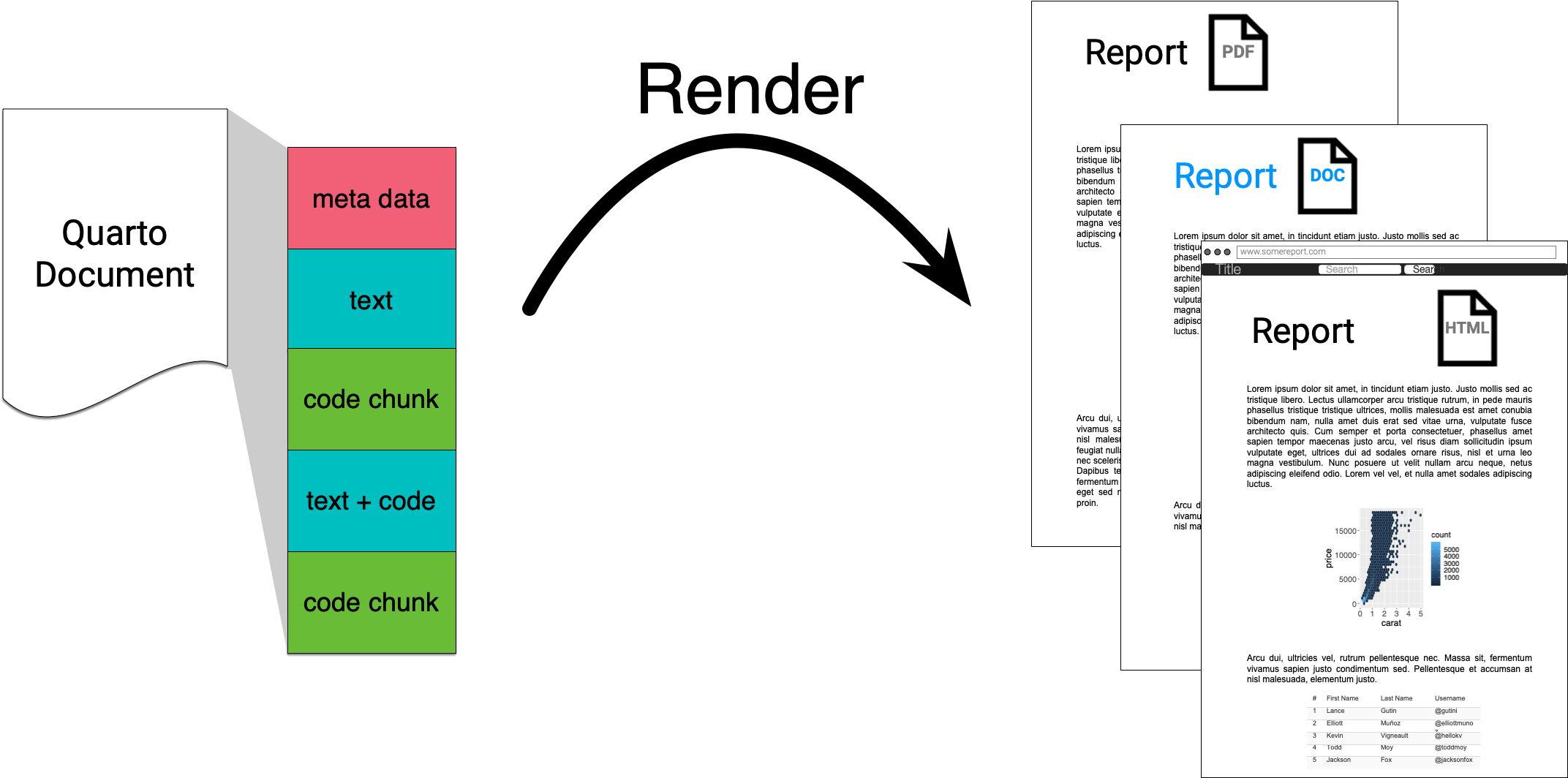
- Note: Quarto is the next generation of R Markdown.
R Markdown
- Quarto and R Markdown are very similar.
- The same team that created R Markdown created Quarto.
- Quarto supersedes R Markdown so we focus on Quarto.
R Markdown
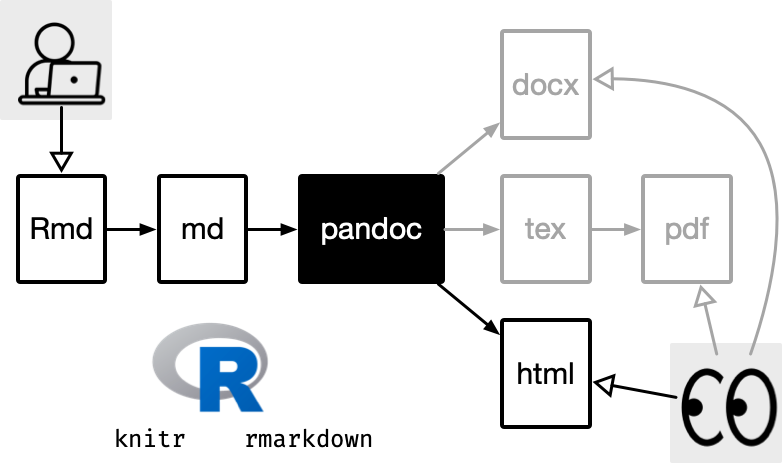
Quarto
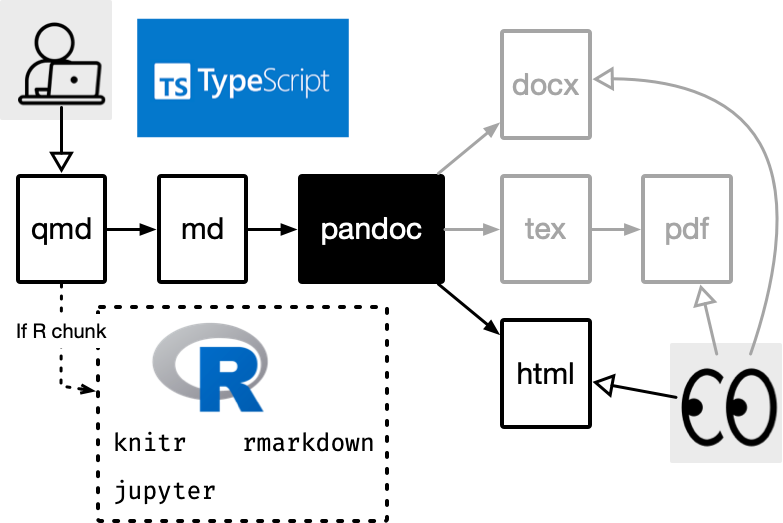
What can you do with Quarto?
There are so many possible output formats you can create with Quarto, including but not limited to:
- Microsoft Word document (.doc, .docx)
- PowerPoint presentation (.pptx)
- Open Document Text (.odt)
- Rich text format (.rtf)
- e-book format (.epub)
- Markdown documents (.md)
- Dashboard (.html)
- Books (.pdf or .html)
- Webpage (.html)
- Websites (collection of web pages)
Primary languages supported:
- R
- Python
- Julia
- Observable
But include engines for many more languages!
HTML slides
These HTML slides are made using Quarto.
Report
These dynamic reports are made using Quarto.
Thesis
This PhD thesis (online and pdf) is made using Quarto.
Available at https://thesis.patrickli.org/
Quarto basics
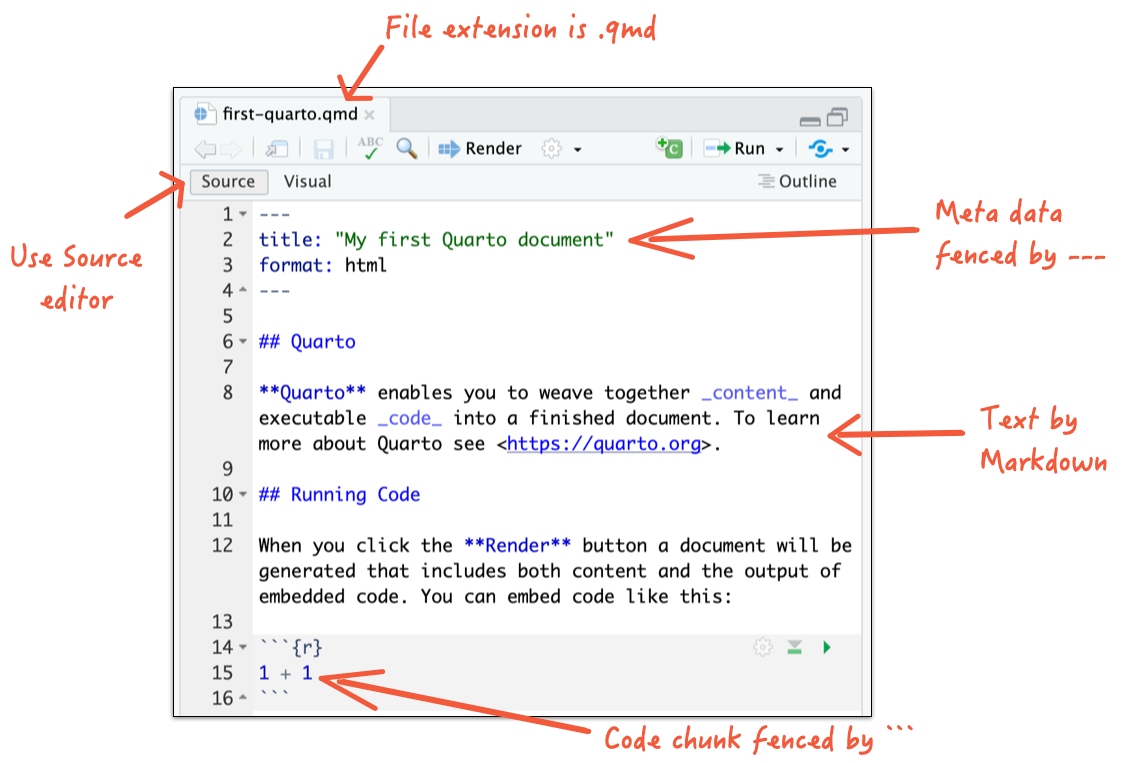
- If you are not using RStudio Desktop, open with your own editor.
Render to a HTML document
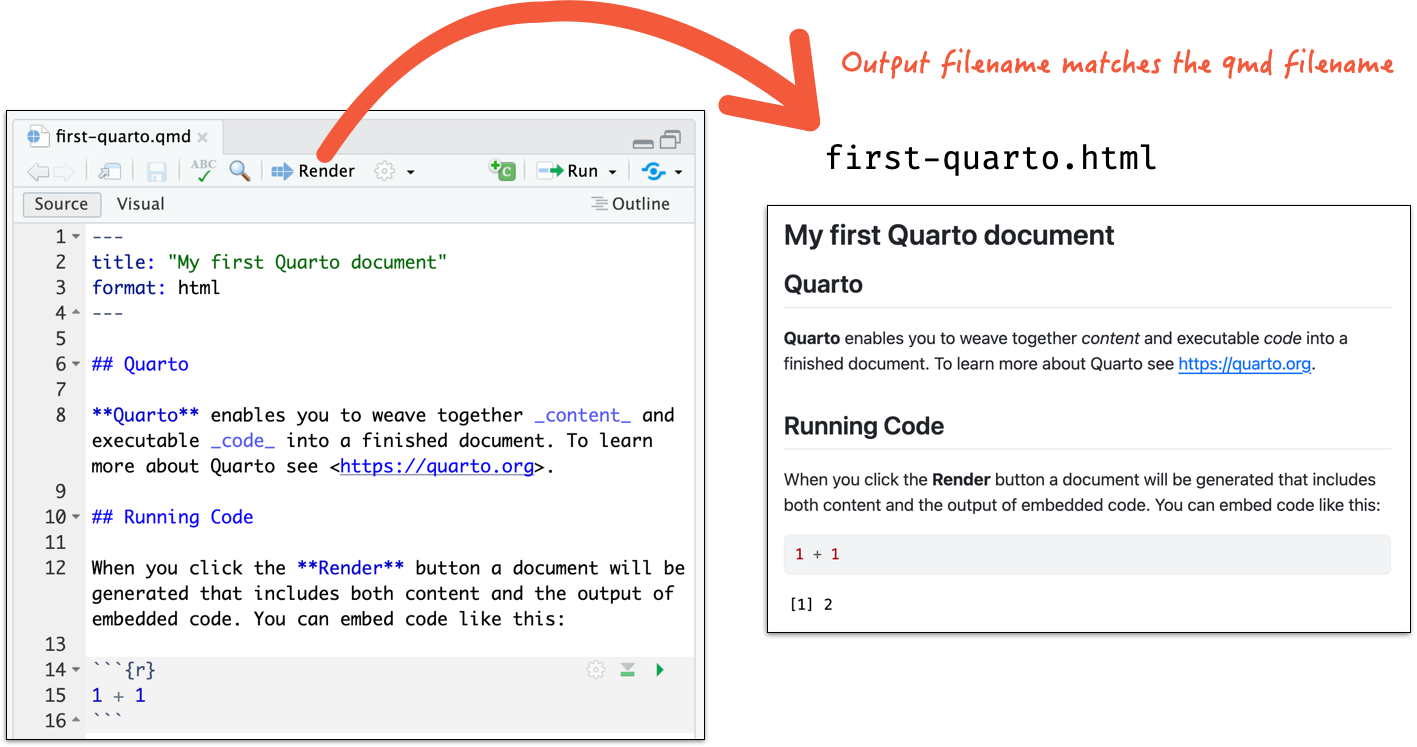
- If you are not using RStudio Desktop, open the terminal and run
Render to a PDF document
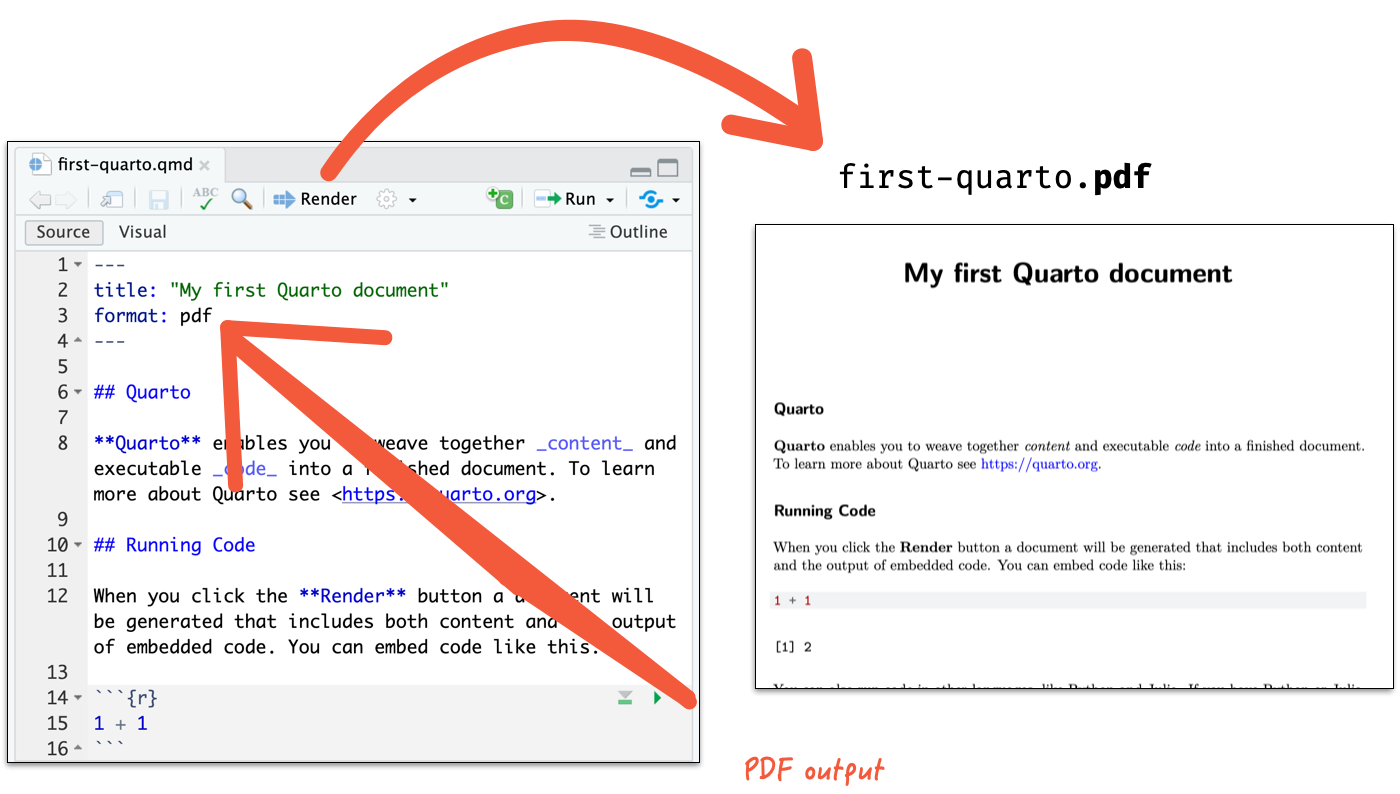
- If you are not using RStudio Desktop, open the terminal and run
RStudio Desktop
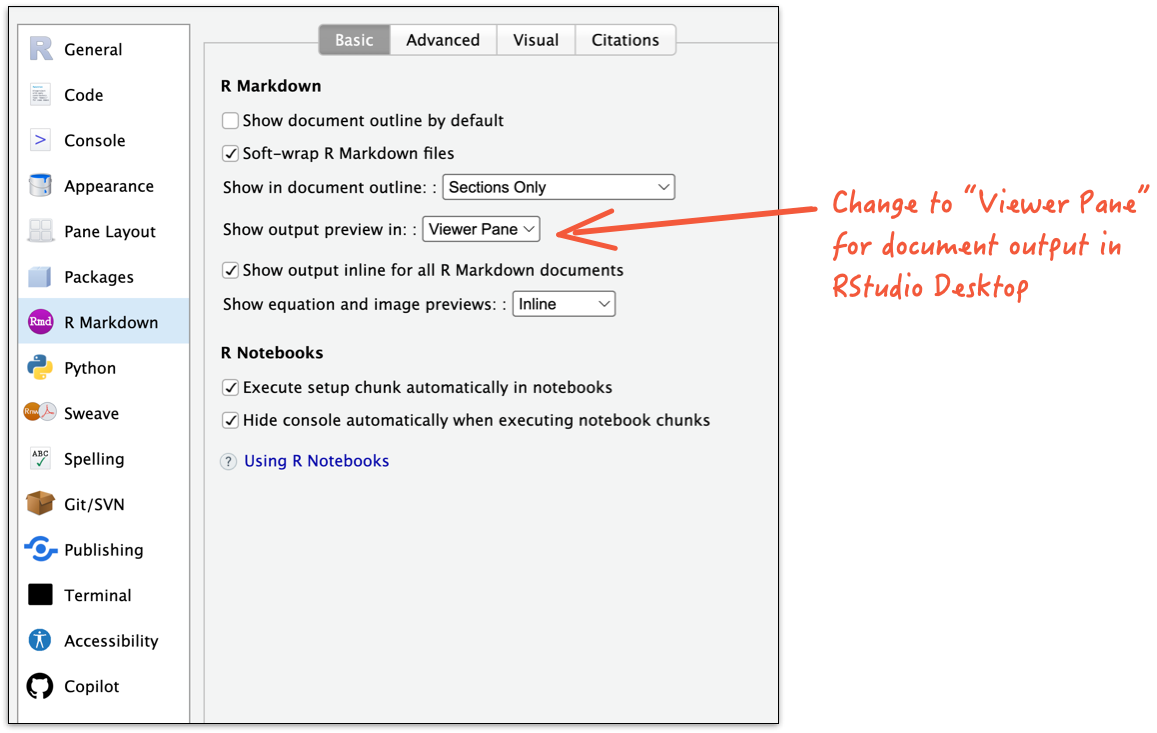
How does it all work?
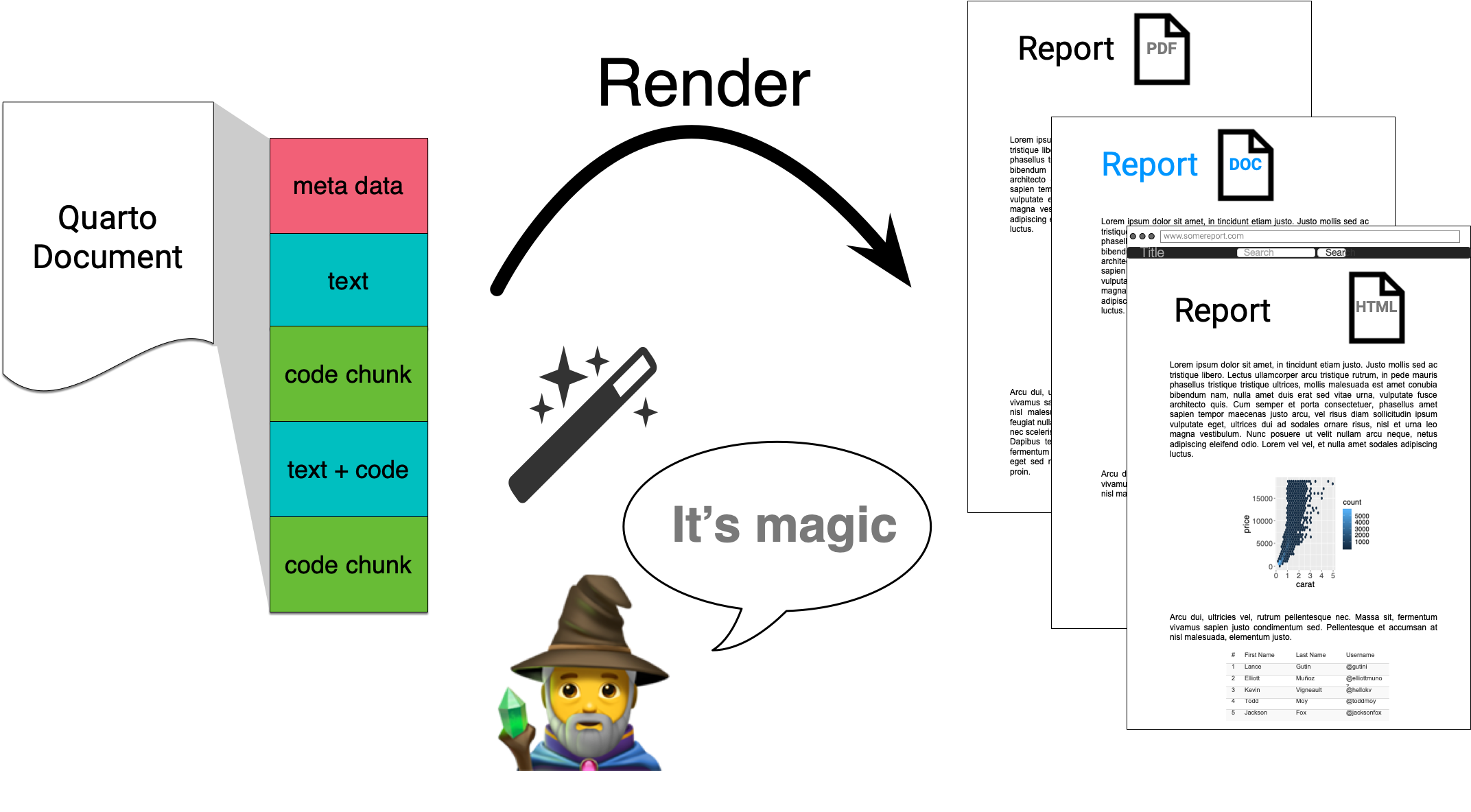
Quarto via knitr/jupyter: qmd md
Pandoc: md html, pdf, docx
Meta Data with YAML
YAML - YAML Ain’t Markup Language
- There must be a space after “
:”! - White spaces indicate structure in YAML - don’t use tabs though!
- Same as R, you can comment lines by starting with
#. - YAML is case sensitive.
- Logical values are specified as
trueorfalse(all lowercase) in YAML (notTRUEorFALSElike in R). - If the value is a string and contains special characters (e.g.,
:,#,-), it should be enclosed in quotes.
YAML in Quarto documents
In Quarto documents, YAML metadata is usually placed at the very top of the document, enclosed by triple dashes ---.
- Some common keys used in Quarto documents:
title- the title of the documentsubtitle- the subtitle of the documentauthor- the author of the documentdate- the date of the documentabstract- a brief summary of the documentformat- the output format of the document (e.g.,html,pdf,docx)
You can find available keys by format at https://quarto.org/docs/reference/
YAML with multiple key values and nested keys
Values spanning multiple lines
A value can span multiple lines in two ways:
- Using the pipe symbol
|to preserve line breaks. - Using the greater-than symbol
>to fold lines (line breaks become spaces).
Text with Markdown
Headers
Lists
Formatting text
Inserting images and links
RStudio > Help > Markdown Quick Reference
Dynamic Documents with Data Analysis Code
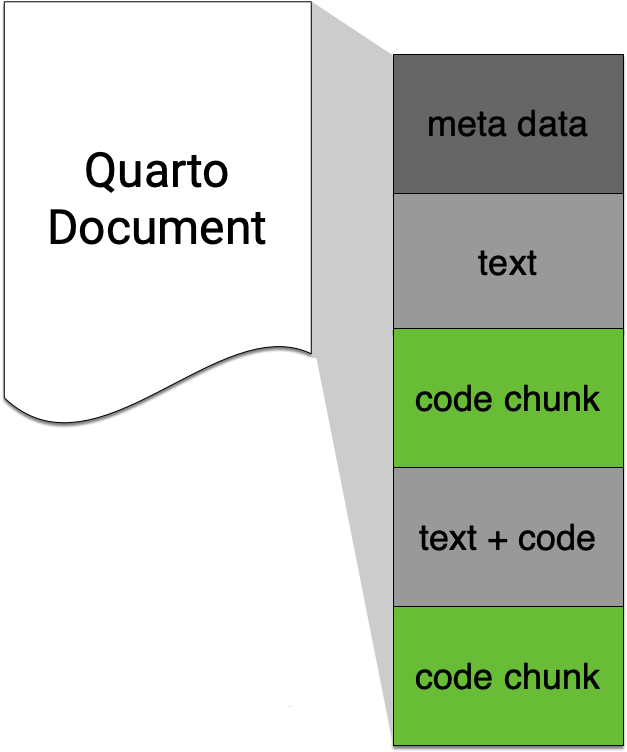
Code chunks
- In Quarto (and R Markdown), code is included in code chunks.
- Code chunks are delimited by triple backticks
```with the language specified after the opening backticks.
Chunk options
label: label for the chunk (for cross-referencing)eval: whether to evaluate the code (trueorfalse)echo: whether to show the code in the output (trueorfalse)fig-width: width of the figure (in inches)fig-height: height of the figure (in inches)fig-cap: caption for the figure
See more options for the knitr engine at here.
Quarto (and R Markdown) is not just for R
- To use Python, change the language to
python:
The following languages are supported by knitr:
asis, asy, awk, bash, block, block2, bslib, c, cat, cc, coffee, comment, css, ditaa, dot, embed, eviews, exec, fortran, fortran95, gawk, glue, glue_sql, gluesql, go, groovy, haskell, highlight, js, julia, lein, mermaid, mysql, node, octave, ojs, perl, php, psql, python, r, rcpp, rscript, ruby, sas, sass, scala, scss, sed, sh, sql, stan, stata, targets, tikz, verbatim, webr, zsh
Inline R code
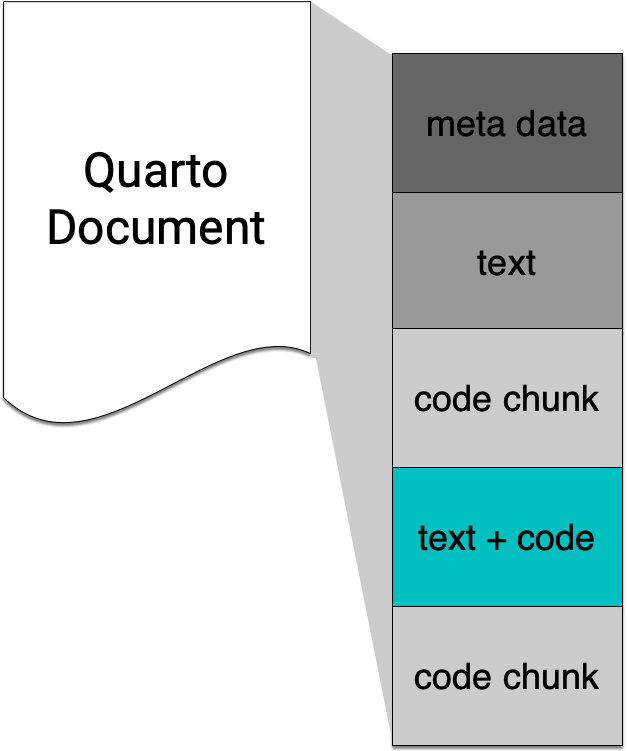
`r some_r_code()`The number of observations in the ChickWeight dataset is 578.
The value of \(\pi\) is 3.1415927.
- Note that these inline R command only work if
engine: knitr. - This doesn’t work for other languages.
Cross Reference
Bibliography
- BibTeX citation style format is used to store references in
.bibfiles. - You can get most BibTeX citation for R packages
citationfunction.
To cite ggplot2 in publications, please use
H. Wickham. ggplot2: Elegant Graphics for Data
Analysis. Springer-Verlag New York, 2016.
A BibTeX entry for LaTeX users is
@Book{,
author = {Hadley Wickham},
title = {ggplot2: Elegant Graphics for Data Analysis},
publisher = {Springer-Verlag New York},
year = {2016},
isbn = {978-3-319-24277-4},
url = {https://ggplot2.tidyverse.org},
}Citing literature
You can cite references like:
- Data analysis was conducted using R (R Core Team 2025)
- Wickham (2016) was used for plotting.
References
- You can cite references in the text using
[@key]or@key, wherekeyis the citation key defined in the.bibfile. - References are automatically appended at the end of the document.
ref.bib
@Book{ggplot2,
author = {Hadley Wickham},
title = {ggplot2: Elegant Graphics for Data Analysis},
publisher = {Springer-Verlag New York},
year = {2016},
isbn = {978-3-319-24277-4},
url = {https://ggplot2.tidyverse.org},
}
@Manual{rstats,
title = {R: A Language and Environment for Statistical Computing},
author = {{R Core Team}},
organization = {R Foundation for Statistical Computing},
address = {Vienna, Austria},
year = {2025},
url = {https://www.R-project.org/},
}Figure references
The chunk label with prefix fig- can be referenced in text as a figure.
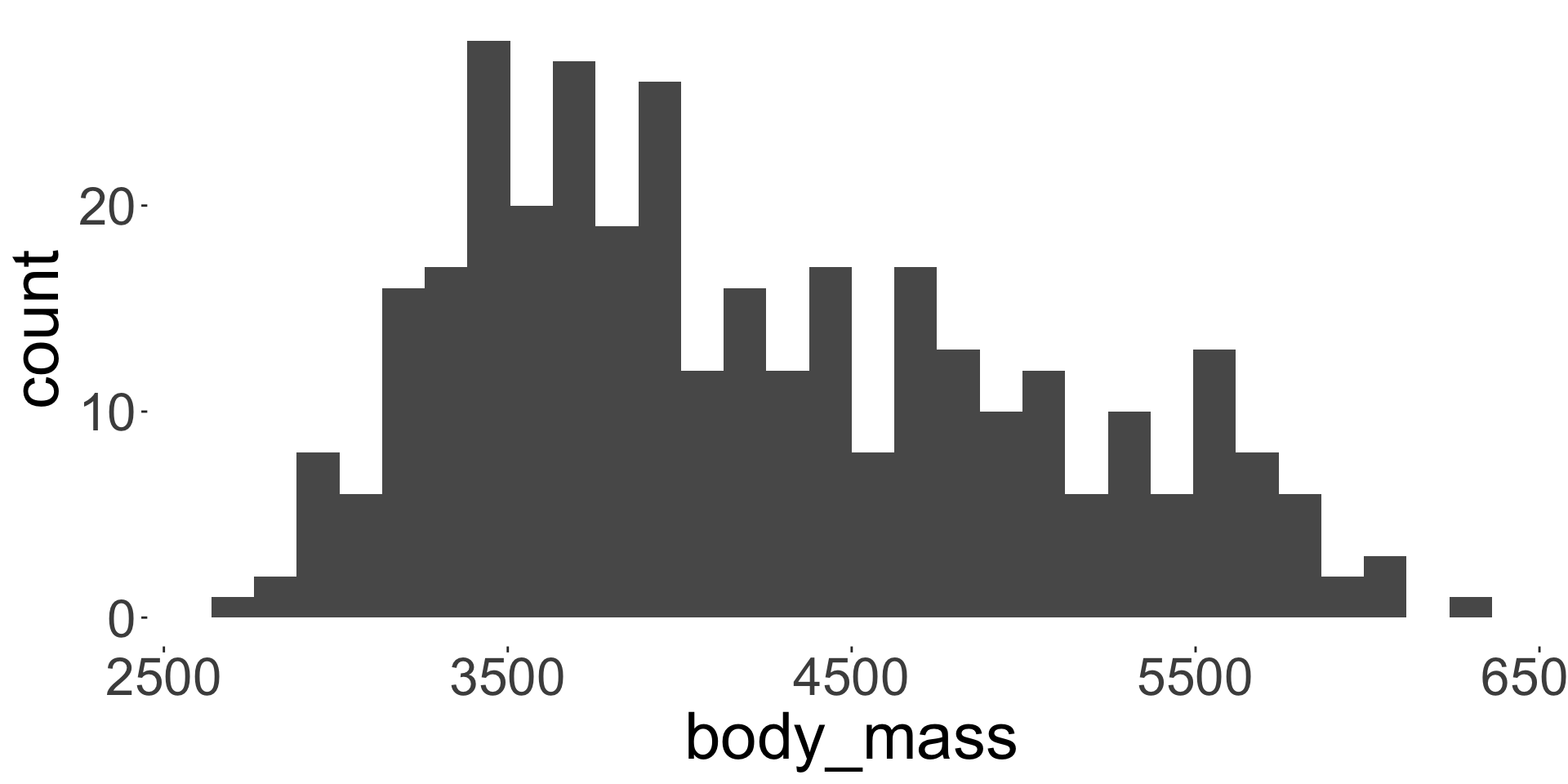
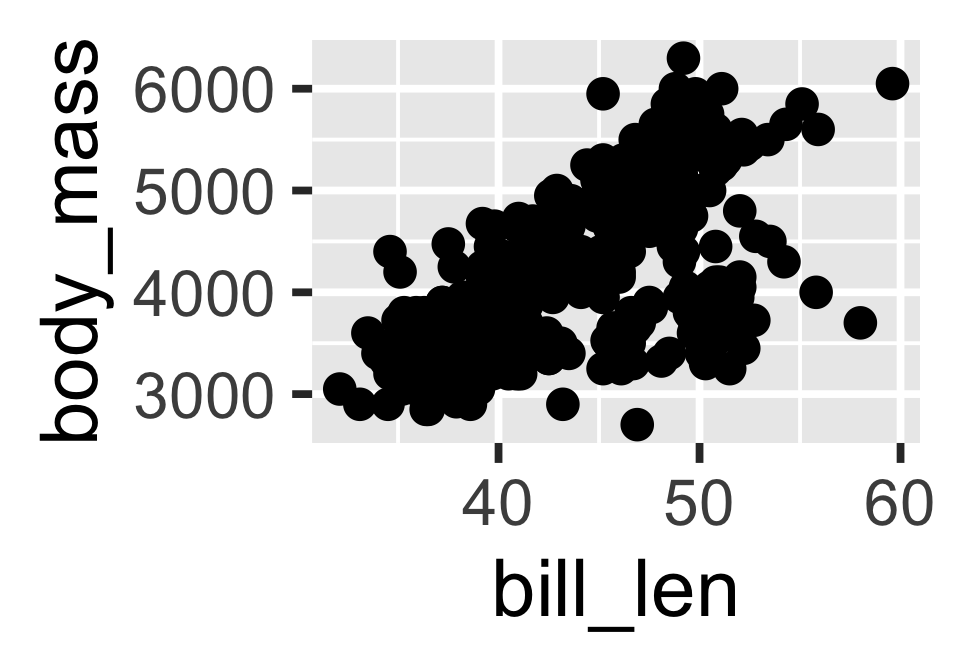
The body mass distribution of penguins is shown in Figure 2.
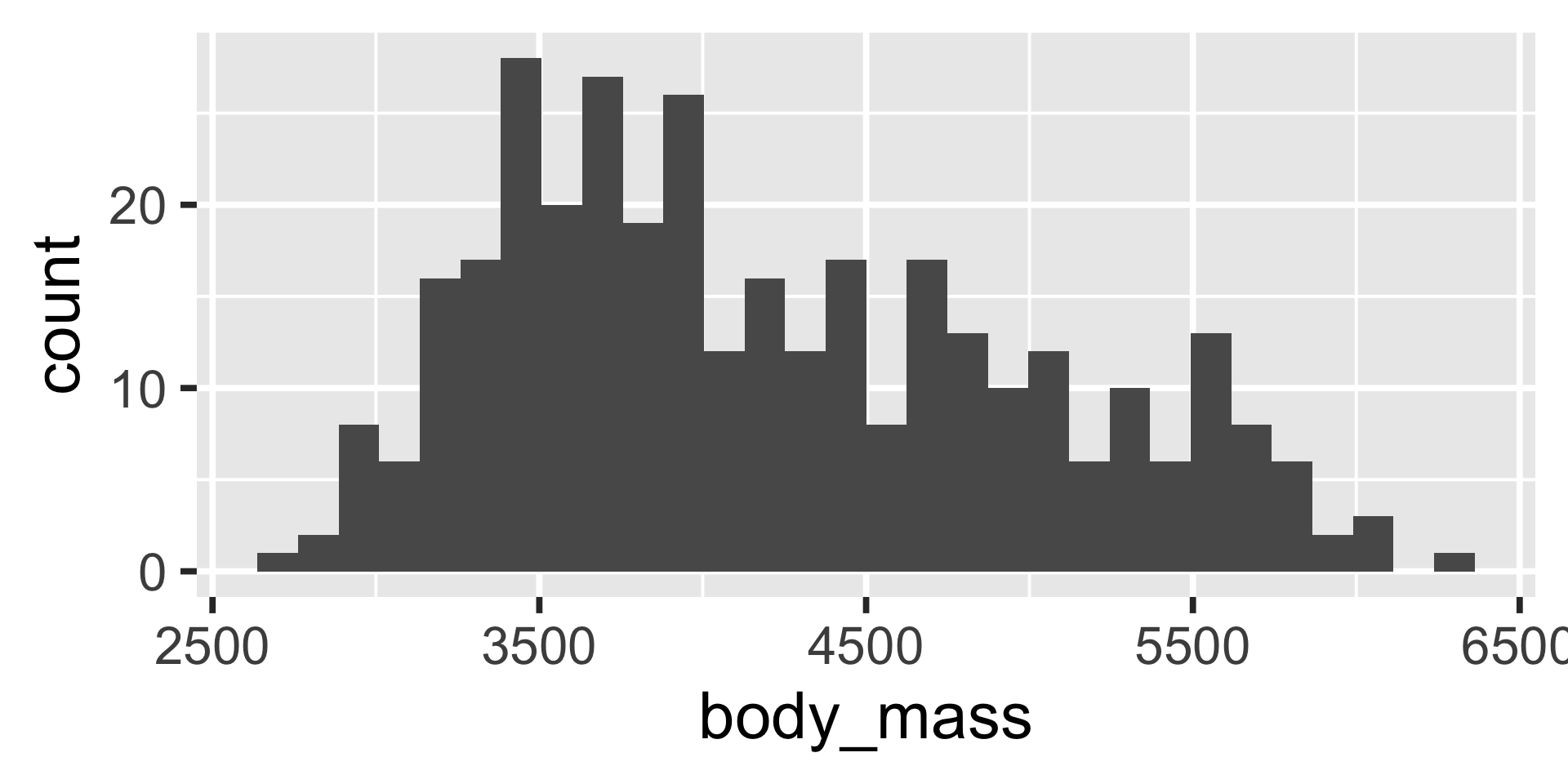
- Above we use the
ggplot2package to create a figure, but you can use base R plotting functions or other packages.
Table references
The chunk label with prefix tbl- can be referenced in text as a table.
| species | Mean | SD | N |
|---|---|---|---|
| Adelie | 3700.7 | 458.6 | 152 |
| Gentoo | 5076.0 | 504.1 | 124 |
| Chinstrap | 3733.1 | 384.3 | 68 |
- Above we use the
knitr::kable()function to create a table. - There are many other packages to create tables, e.g.,
gt,flextable,kableExtra, etc.
Section references
If you have enabled numbering of sections:
then you can refer to them by their label prefixed by sec-.
Summary

Weave together text, code, and output (figures, tables, etc.) in a single document using Quarto into various output formats (HTML, PDF, Word, etc.).
- Use YAML to control document’s meta data.
- Use Markdown syntax in the body of the document to write content.
- Use R (or Python, Julia, etc.) for data analysis and visualization.
- Focus on writing the content instead of formatting!
- The best guide for Quarto is at https://quarto.org/docs/guide/.
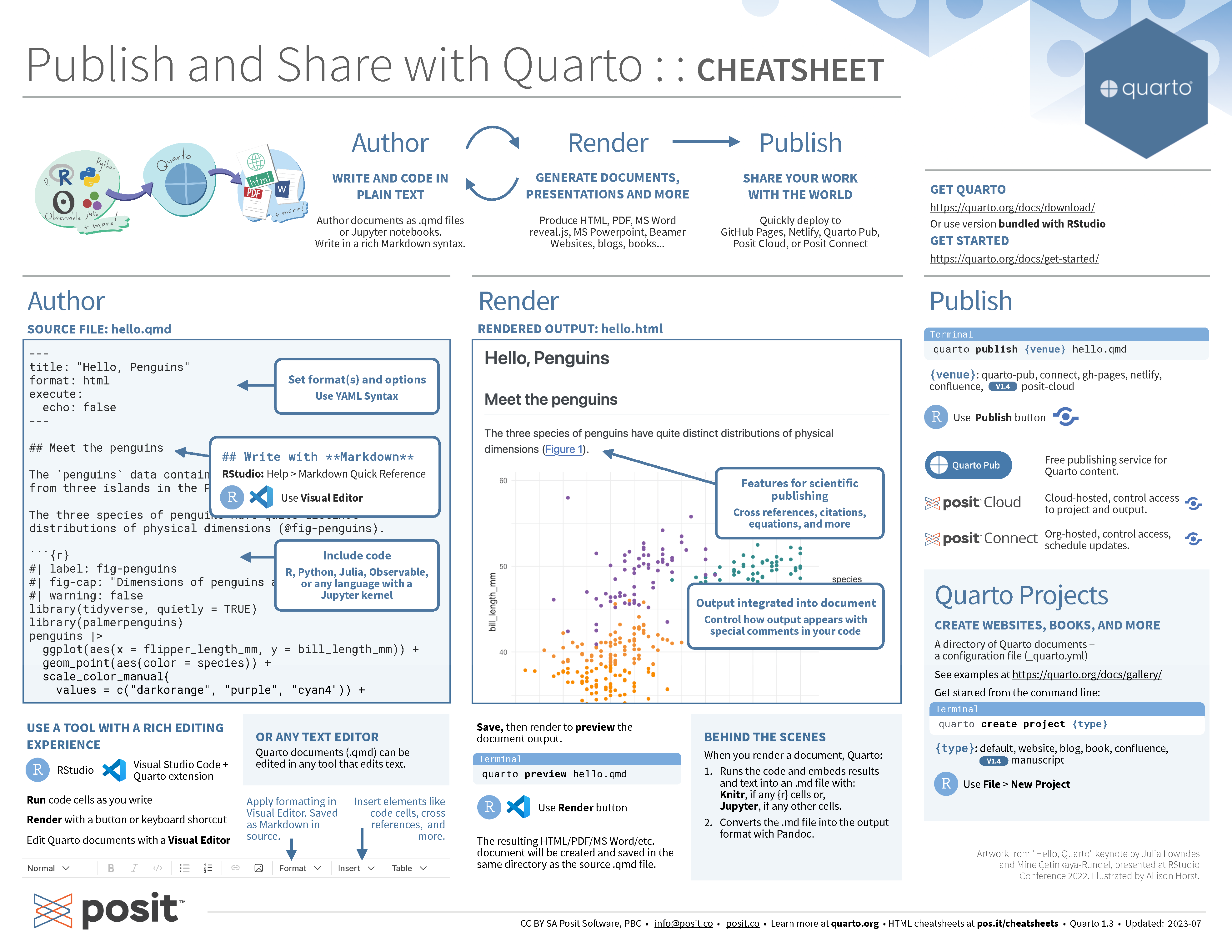
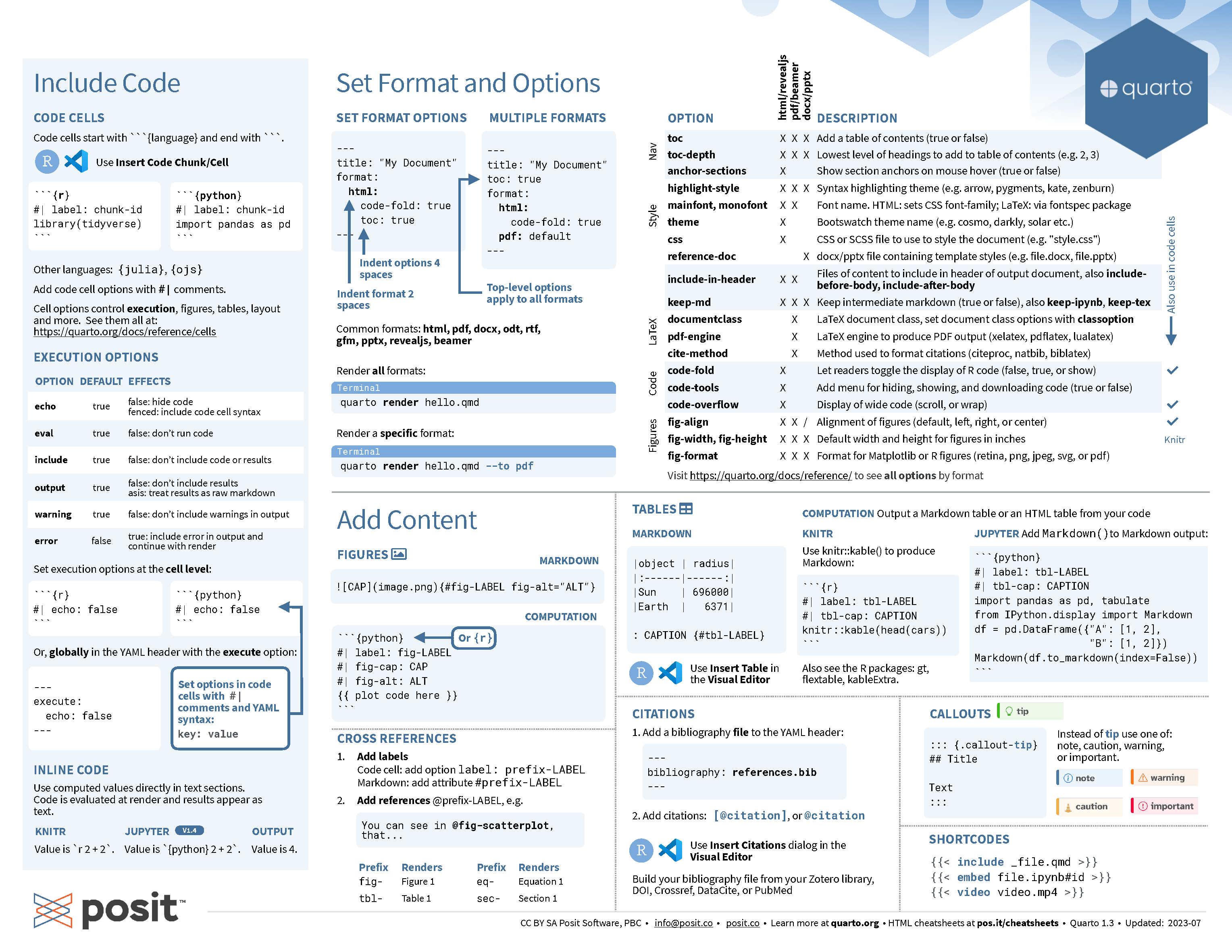
Quarto cheatsheet


Literate Programming
Non-robust workflow
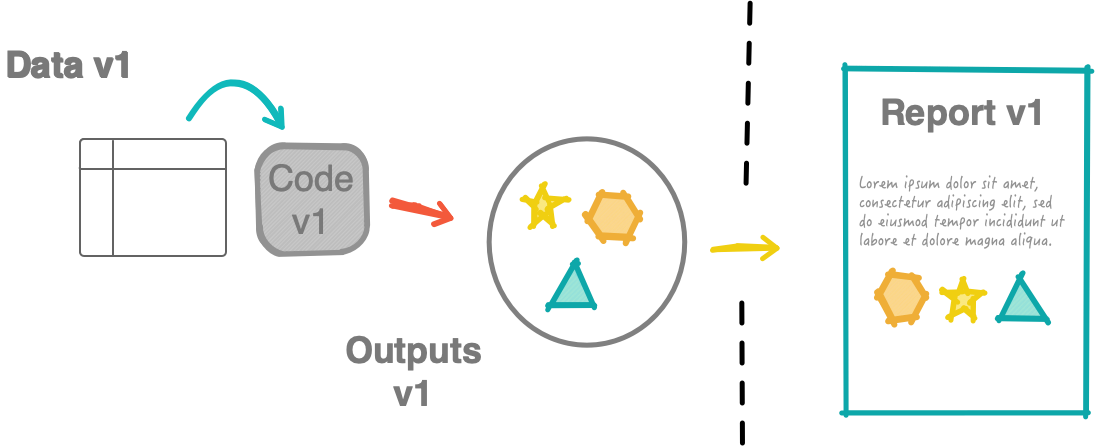
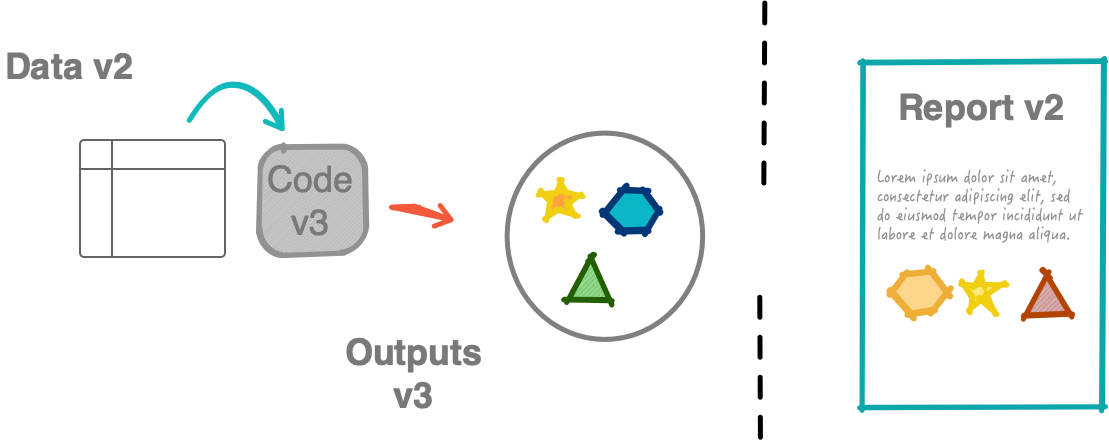
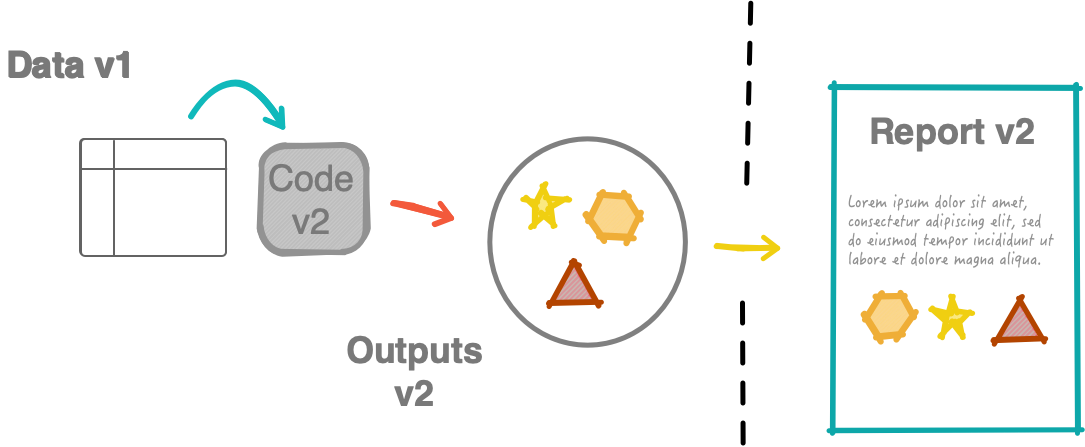
What should have been submitted:

A robust, reproducible workflow
- Using a robust, reproducible workflow means:
- you avoid manual, repetitive tasks
- your results are computationally reproducible
- Using a robust, reproducible workflow doesn’t mean you won’t make mistakes, but it will help you minimise mistakes.
Literate programming is a programming paradigm introduced by Donald Knuth where it emphasises writing code for humans (i.e. intertwine code with natural language explanations).
- Literate programming includes documentation (detailed explanations, comments and annotations to provide context, rationale and insight into the program’s design and functionality).
Analysis framework
Tidy data
- Each column is a variable.
- Each row is an observation.
- Each cell is a single value.
Tools
- Git/GitHub for version control and collaboration
- Open-source programming languages (e.g. R and Python) for coding
- Quarto with markdown syntax for interoperable reproducible reports
Statistical value chain
… a statistical value chain is constructed by defining a number of meaningful intermediate data products, for which a chosen set of quality attributes are well described …
– van der Loo & de Jonge (2018)
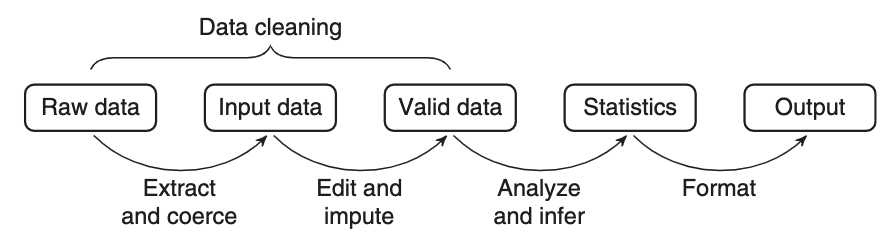
Folder structure
A suggested folder structure for data projects:
project-root-folder/ # Root of the project folder
│
├── README.md # README file
│
├── data/ # Raw and derived data
│ ├── data-raw/ # Read-only files
│ ├── data-input/ # Extracted and coerced from raw data
│ ├── data-valid/ # Edit and imputed from input data
│ └── data-stats/ # Analysed results (R objects, .csv, etc.)
│
├── analysis/ # Scripts (not functions) to run analysis
│
├── figures/ # Figures (.png, .pdf, etc.)
│
├── misc/ # Misc
│
├── report.qmd # Report, paper, or thesis outputSharing your documents
via Quarto Pubs
- Make sure you are logged in to your Quarto Pub account.
- Then run the following command in the Terminal:
Happy writing and sharing 😊

STAT1003 – Statistical Techniques