Joining two datasets
STAT1003 – Statistical Techniques
Dr. Emi Tanaka
Australian National University
These slides are best viewed on a modern browser like Google Chrome on a desktop or laptop. Some interactive components may require some time to fully load.
Relational model
- A relational model organizes data into one or more tables of columns and rows, with a unique key identifying each row based on Codd (1969, 1970).
- Generally, each table represents one entity type.
- Relational database is a database based on the relational model of data.
| Type | Tidy data | Relational model |
|---|---|---|
| variable | column | attribute/field |
| observation | row | tuple/record |
Primary key
- A primary key (sometimes called candidate key) is the smallest subset of columns that uniquely identifies each row in a table. E.g.
species_idis the primary key in thespeciestable,plot_idis the primary key in theplotstable, andrecord_idis the primary key in thesurveystable.
Simple key & Compound key
- If only a single column then it is called a simple key.
- If a key consists of more than one column then it is called a compound key.
- E.g.
ChickWeighthas a compound key consisting ofTimeandChick.
- E.g.
- A table can also have no key (violating the relational model).
- E.g.
chickwtshas no key.
- E.g.
Foreign key
- A foreign key is a column in one table that uniquely identifies a row in another table.
plot_idandspecies_idare foreign keys in thesurveystable, which link to theplotsandspeciestables, respectively.- There are no foreign keys in the
plotsandspeciestables.
Joining tables
- To join tables, we use the primary key in one table and the foreign key in another table.
- In a relational model, a database has referential integrity if all relations between tables are valid. E.g.,
- All primary key values must be unique and not missing.
- Each foreign key value must have a corresponding primary key value.
- In a relational model, normalization aims to keep data organization as clean and simple as possible by avoiding redundant data entries.
Equality (equi) joins in dplyr
- Natural join is a special case of equality join where the join is performed on all columns with the same name in both tables.
- In
dplyr, you can useby = NULL(default) to perform a natural join.
Cardinality in data modelling
- The relationship between the two tables can be categorized into four types:
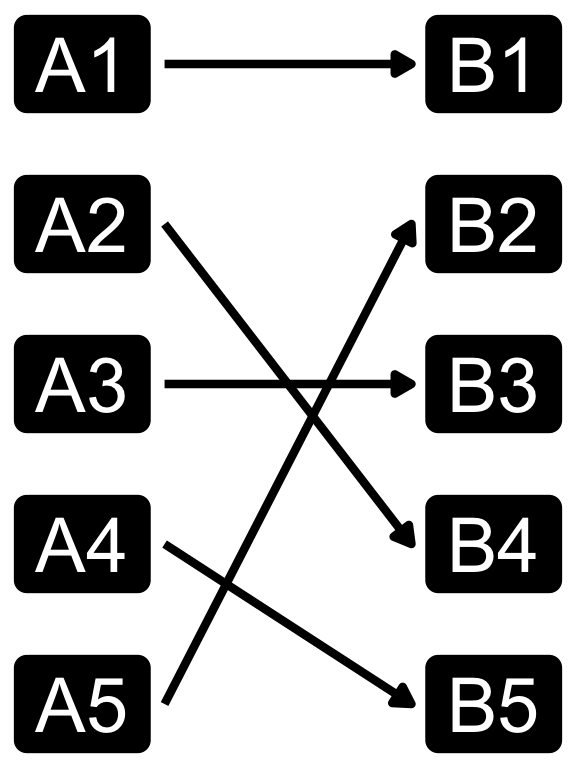
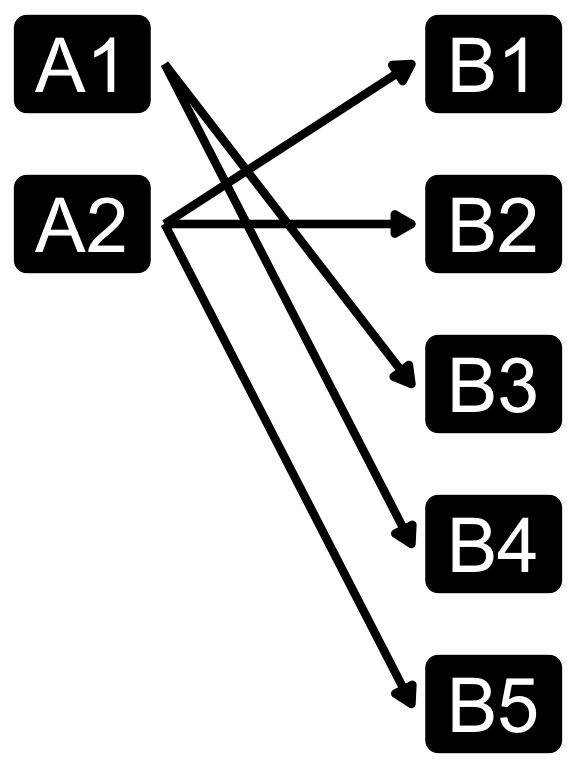
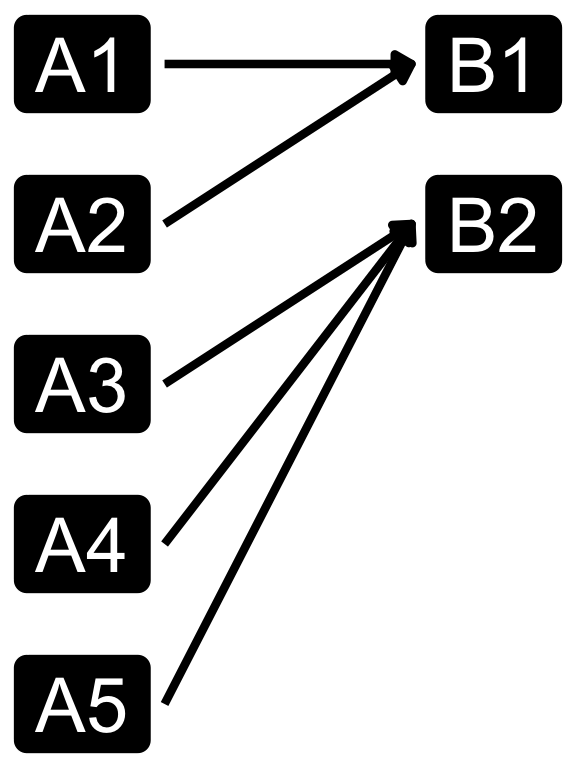
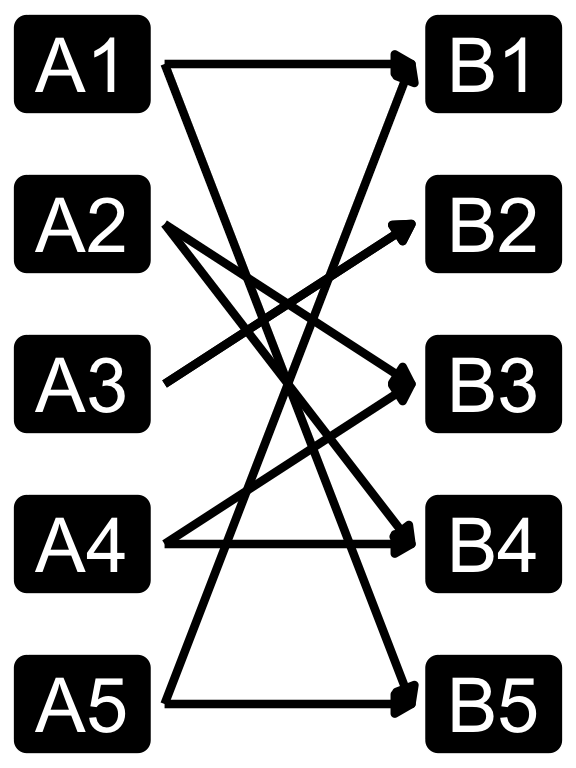
- You can specify relationshp in
dplyrjoins using therelationshipargument, which can be set to “one-to-one”, “one-to-many”, “many-to-one”, or “many-to-many”. - Note that relationship is not a property of the data, but rather of the relationship itself.
Challenge 2022 election result in the ACT
| name | enrol |
|---|---|
| BEAN | 109557 |
| CANBERRA | 102196 |
| FENNER | 102576 |
| division | party | votes |
|---|---|---|
| Bean | AUP | 2722 |
| Bean | LP | 29223 |
| Bean | GAP | 929 |
| Bean | LDP | 2540 |
| Bean | GRN | 12168 |
| Bean | UAPP | 2227 |
| Bean | ALP | 35447 |
| Bean | 5043 | |
| Bean | IND | 7683 |
| Canberra | AUP | 1784 |
| Canberra | GRN | 20144 |
| Canberra | 1904 | |
| Canberra | UAPP | 1361 |
| Canberra | LP | 24063 |
| Canberra | ALP | 34989 |
| Canberra | IND | 4062 |
| Fenner | 2669 | |
| Fenner | ALP | 38864 |
| Fenner | UAPP | 3529 |
| Fenner | GRN | 12492 |
| Fenner | LP | 30025 |
| Fenner | AUP | 1723 |
| division | party | votes | enrol | perc |
|---|---|---|---|---|
| BEAN | AUP | 2722 | 109557 | 2.5 |
| BEAN | LP | 29223 | 109557 | 26.7 |
| BEAN | GAP | 929 | 109557 | 0.8 |
| BEAN | LDP | 2540 | 109557 | 2.3 |
| BEAN | GRN | 12168 | 109557 | 11.1 |
| BEAN | UAPP | 2227 | 109557 | 2.0 |
| BEAN | ALP | 35447 | 109557 | 32.4 |
| BEAN | 5043 | 109557 | 4.6 | |
| BEAN | IND | 7683 | 109557 | 7.0 |
| CANBERRA | AUP | 1784 | 102196 | 1.7 |
| CANBERRA | GRN | 20144 | 102196 | 19.7 |
| CANBERRA | 1904 | 102196 | 1.9 | |
| CANBERRA | UAPP | 1361 | 102196 | 1.3 |
| CANBERRA | LP | 24063 | 102196 | 23.5 |
| CANBERRA | ALP | 34989 | 102196 | 34.2 |
| CANBERRA | IND | 4062 | 102196 | 4.0 |
| FENNER | 2669 | 102576 | 2.6 | |
| FENNER | ALP | 38864 | 102576 | 37.9 |
| FENNER | UAPP | 3529 | 102576 | 3.4 |
| FENNER | GRN | 12492 | 102576 | 12.2 |
| FENNER | LP | 30025 | 102576 | 29.3 |
| FENNER | AUP | 1723 | 102576 | 1.7 |
Inequality (non-equi) joins in dplyr
| id | name | date |
|---|---|---|
| 1 | Alice | 2021-11-01 |
| 1 | Alice | 2022-01-01 |
| 2 | Bob | 2022-02-15 |
| 3 | Charlie | 2022-03-10 |
| patient_id | name | start | end |
|---|---|---|---|
| 1 | Aspirin | 2021-12-01 | 2022-01-31 |
| 1 | Tylenol | 2022-02-01 | 2022-02-28 |
| 2 | Ibuprofen | 2022-02-20 | 2022-03-15 |
| 3 | Amoxicillin | 2022-01-15 | 2022-02-28 |
| 3 | Vitamin C | 2022-04-01 | 2022-04-30 |
Overlap (interval) joins in dplyr
Genomic data often involves intervals and we may want to join two datasets based on whether the intervals overlap.
| id | chr | start | end |
|---|---|---|---|
| 1 | chr1 | 140 | 150 |
| 2 | chr2 | 210 | 240 |
| 3 | chr2 | 380 | 415 |
| 4 | chr1 | 230 | 280 |
| id | chr | start | end |
|---|---|---|---|
| 1 | chr1 | 100 | 150 |
| 2 | chr1 | 200 | 250 |
| 3 | chr2 | 300 | 399 |
| 4 | chr2 | 415 | 450 |
Summary
- Relational data consists of multiple, linked tables.
- Primary keys uniquely identify each record in a table.
- A foreign key is required to join tables together.
- A key can be simple (one column) or compound (multiple columns).
- Cardinality describes the relationship between tables (one-to-one, one-to-many, many-to-one, or many-to-many) which can be used to ensure data integrity.
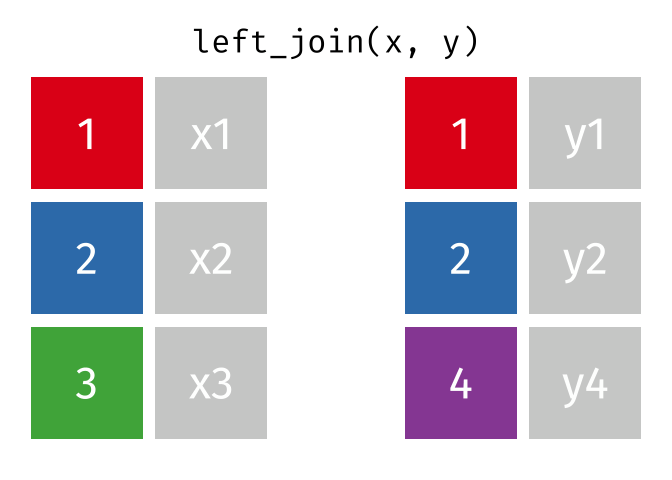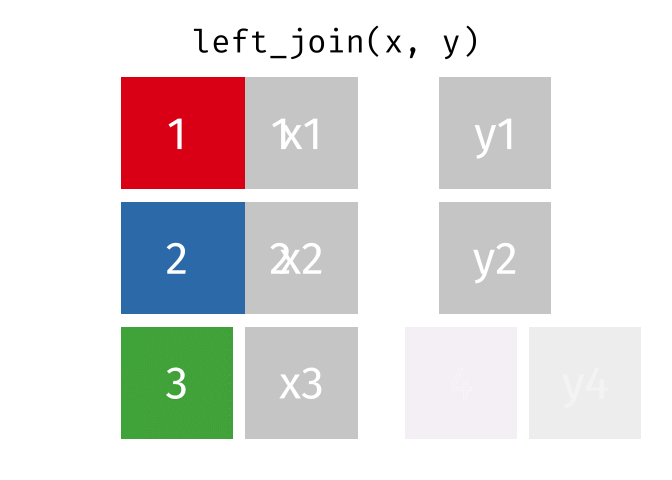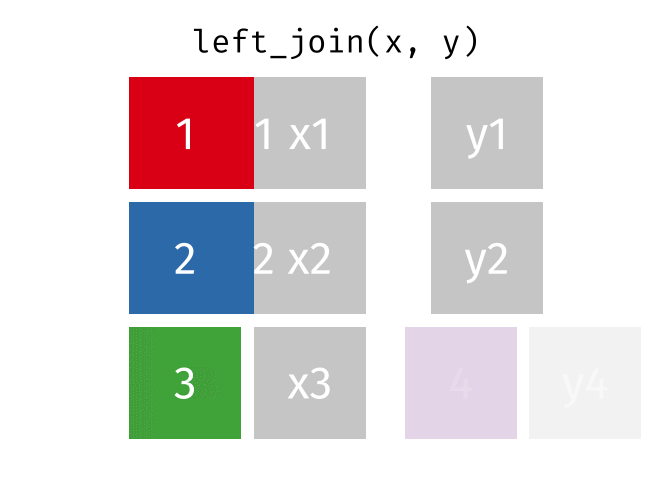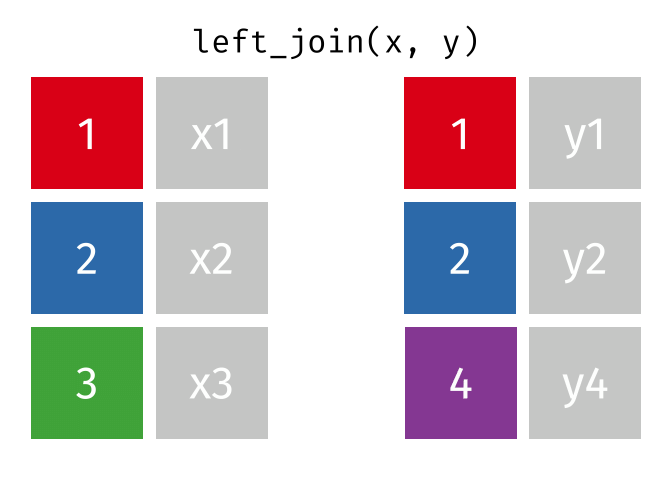
- Left join
- Left join (extra)
- Right join
- Semi join
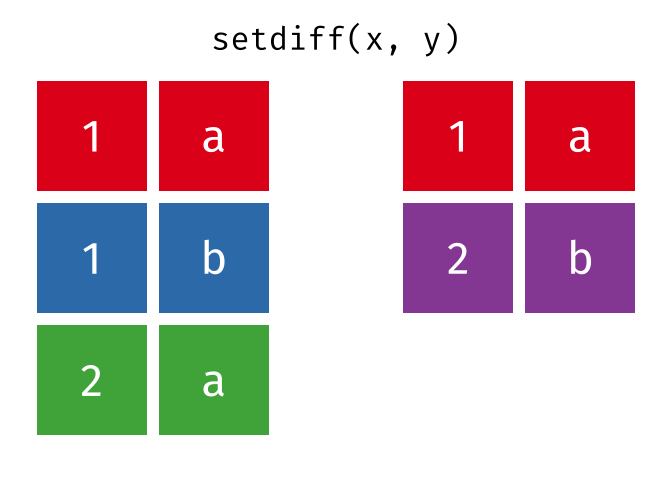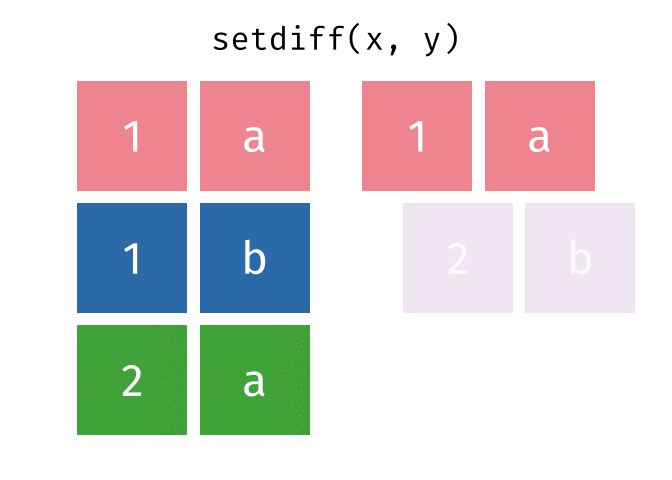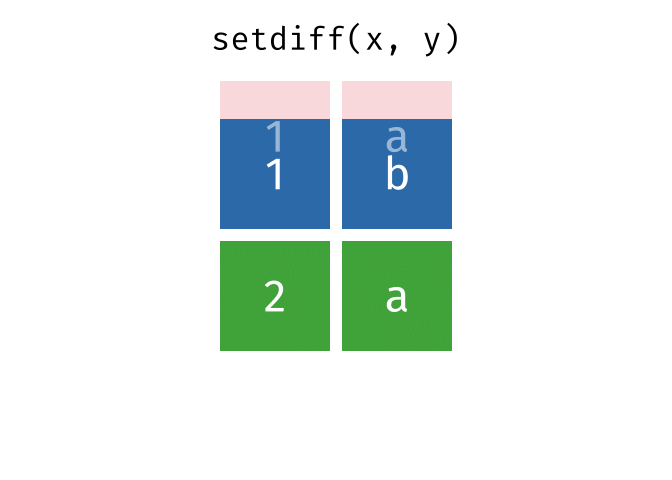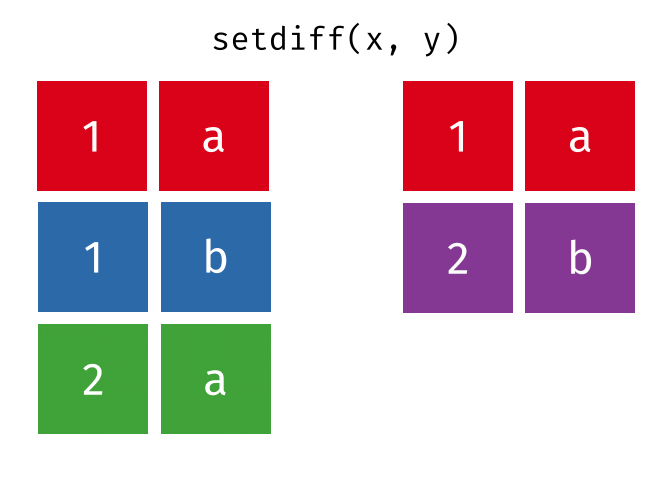
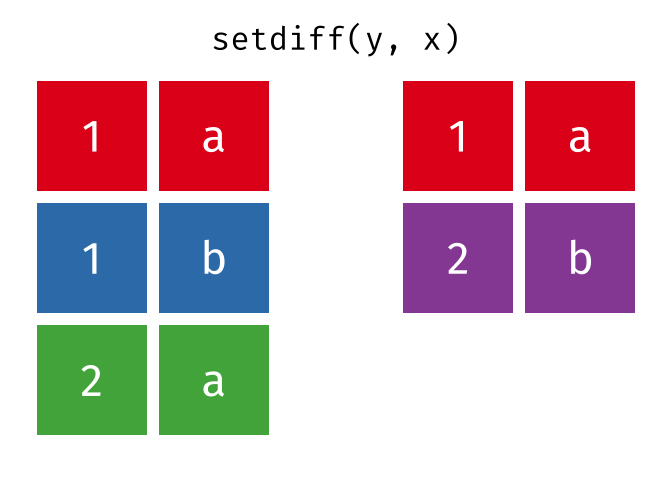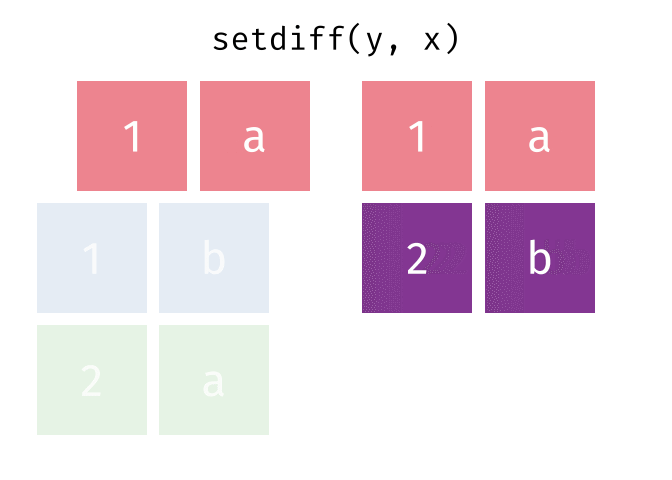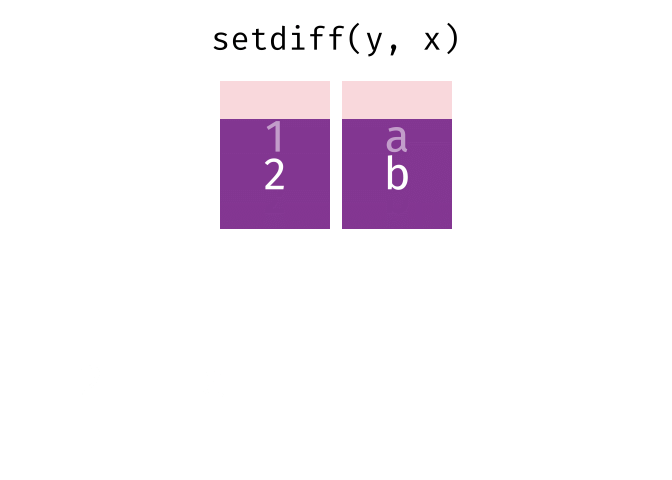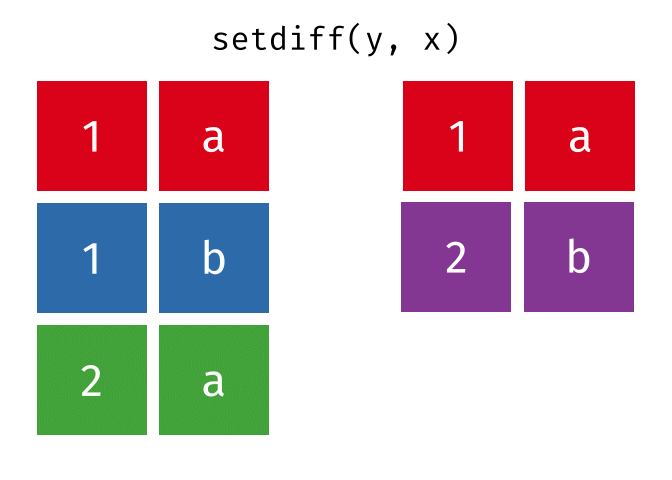
- Set diff
- Set diff (reverse)
- Anti join
- Full join
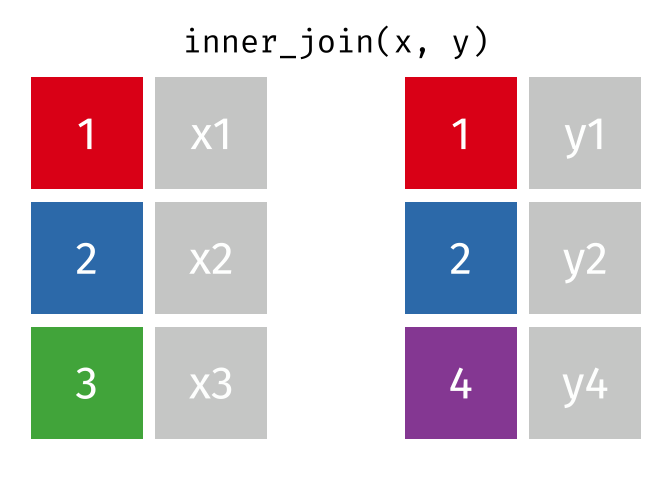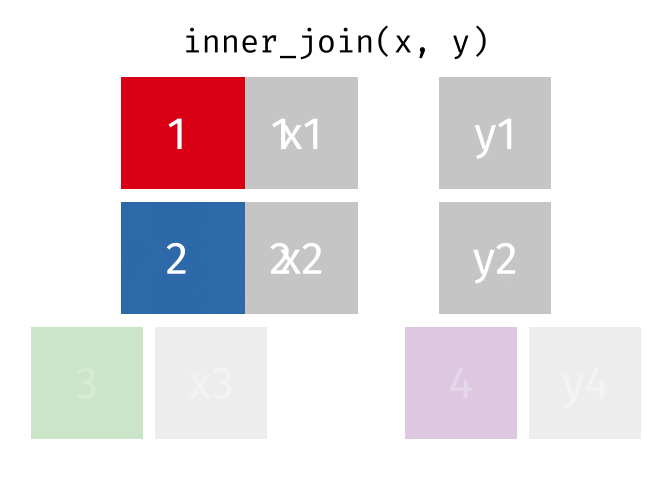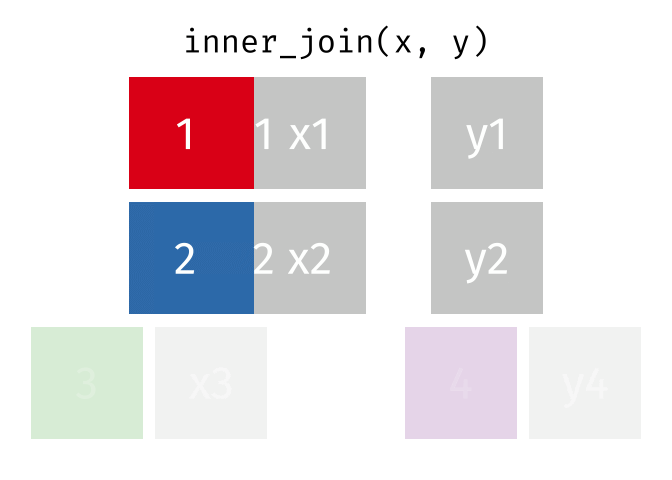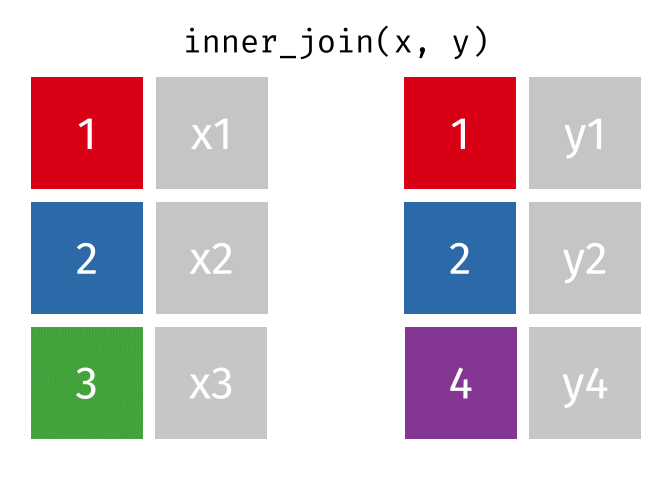
- Inner join
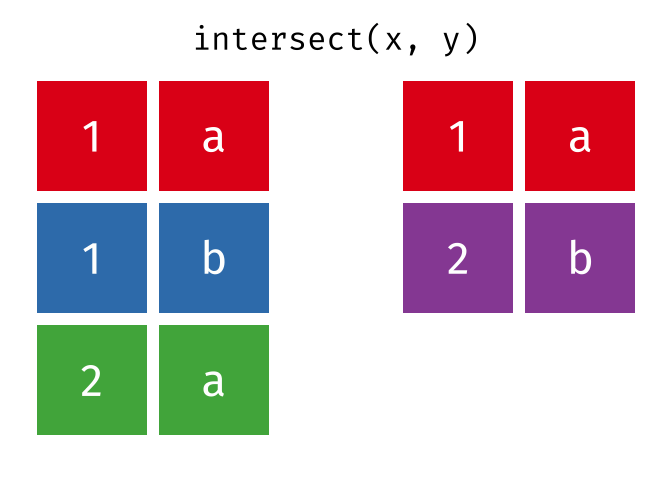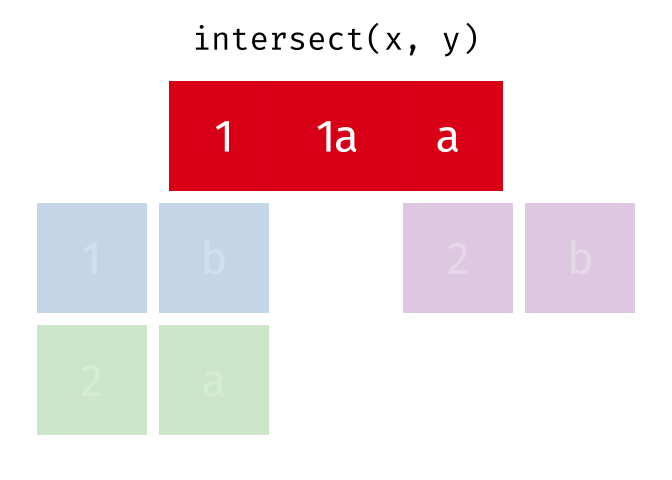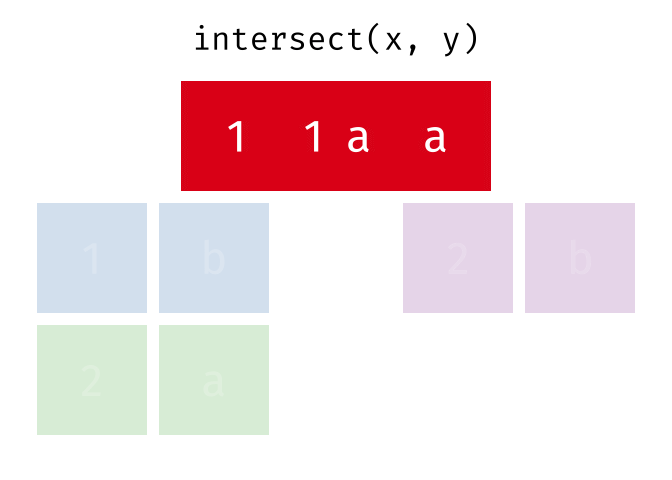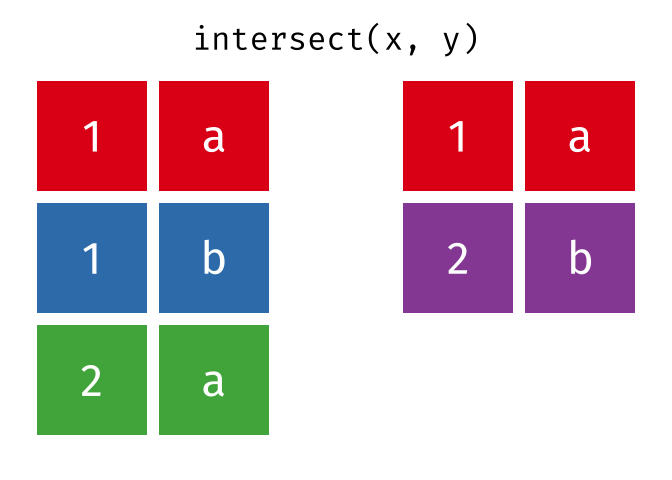
- Intersect






STAT1003 – Statistical Techniques